Computational and mathematical modeling of complex biological systems
 An illustration of the systems approach to biology
An illustration of the systems approach to biology
Systems biology
is the
computational
and
mathematical
analysis and modeling of complex
biological systems
. It is a
biology
-based interdisciplinary field of study that focuses on complex interactions within biological systems, using a holistic approach (
holism
instead of the more traditional
reductionism
) to biological research.
[1]
Particularly from the year 2000 onwards, the concept has been used widely in biology in a variety of contexts. The
Human Genome Project
is an example of applied
systems thinking
in biology which has led to new, collaborative ways of working on problems in the biological field of genetics.
[2]
One of the aims of systems biology is to model and discover
emergent properties
, properties of
cells
,
tissues
and
organisms
functioning as a
system
whose theoretical description is only possible using techniques of systems biology.
[1]
[3]
These typically involve
metabolic networks
or
cell signaling
networks.
[1]
[4]
Overview
[
edit
]
Systems biology can be considered from a number of different aspects.
As a field of study, particularly, the study of the interactions between the components of biological systems, and how these interactions give rise to the function and behavior of that system (for example, the
enzymes
and
metabolites
in a
metabolic pathway
or the heart beats).
[5]
[6]
[7]
As a
paradigm
, systems biology is usually defined in antithesis to the so-called
reductionist
paradigm (
biological organisation
), although it is consistent with the
scientific method
. The distinction between the two paradigms is referred to in these quotations: "the
reductionist
approach has successfully identified most of the components and many of the interactions but, unfortunately, offers no convincing concepts or methods to understand how system properties emerge ... the pluralism of causes and effects in biological networks is better addressed by observing, through quantitative measures, multiple components simultaneously and by rigorous data integration with mathematical models." (Sauer
et al.
)
[8]
"Systems biology ... is about putting together rather than taking apart, integration rather than reduction. It requires that we develop ways of thinking about integration that are as rigorous as our reductionist programmes, but different. ... It means changing our philosophy, in the full sense of the term." (
Denis Noble
)
[7]
As a series of operational
protocols
used for performing research, namely a cycle composed of theory,
analytic
or
computational modelling
to propose specific testable hypotheses about a biological system, experimental validation, and then using the newly acquired quantitative description of cells or cell processes to refine the computational model or theory.
[9]
Since the objective is a model of the interactions in a system, the experimental techniques that most suit systems biology are those that are system-wide and attempt to be as complete as possible. Therefore,
transcriptomics
,
metabolomics
,
proteomics
and
high-throughput techniques
are used to collect quantitative data for the construction and validation of models.
[10]
As the application of
dynamical systems theory
to
molecular biology
. Indeed, the focus on the dynamics of the studied systems is the main conceptual difference between systems biology and
bioinformatics
.
[11]
As a socioscientific phenomenon defined by the strategy of pursuing integration of complex data about the interactions in biological systems from diverse experimental sources using interdisciplinary tools and personnel.
[12]
History
[
edit
]
Although the concept of a systems view of cellular function has been well understood since at least the 1930s,
[13]
technological limitations made it difficult to make systems wide measurements. The advent of microarray technology in the 1990s opened up an entire new visa for studying cells at the systems level. In 2000, the Institute for Systems Biology was established in Seattle in an effort to lure "computational" type people who it was felt were not attracted to the academic settings of the university. The institute did not have a clear definition of what the field actually was: roughly bringing together people from diverse fields to use computers to holistically study biology in new ways.
[14]
A Department of Systems Biology at Harvard Medical School was launched in 2003.
[15]
In 2006 it was predicted that the buzz generated by the "very fashionable" new concept would cause all the major universities to need a systems biology department, thus that there would be careers available for graduates with a modicum of ability in computer programming and biology.
[14]
In 2006 the
National Science Foundation
put forward a challenge to build a mathematical model of the whole cell.
[
citation needed
]
In 2012 the first whole-cell model of
Mycoplasma genitalium
was achieved by the Karr Laboratory at the Mount Sinai School of Medicine in New York. The whole-cell model is able to predict viability of
M. genitalium
cells in response to genetic mutations.
[16]
An earlier precursor of systems biology, as a distinct discipline, may have been by systems theorist
Mihajlo Mesarovic
in 1966 with an international symposium at the
Case Institute of Technology
in
Cleveland
, Ohio, titled
Systems Theory and Biology
. Mesarovic predicted that perhaps in the future there would be such a thing as "systems biology".
[17]
[18]
Other early precursors that focused on the view that biology should be analyzed as a system, rather than a simple collection of parts, were
Metabolic Control Analysis
, developed by
Henrik Kacser
and Jim Burns
[19]
later thoroughly revised,
[20]
and
Reinhart Heinrich
and
Tom Rapoport
,
[21]
and
Biochemical Systems Theory
developed by
Michael Savageau
[22]
[23]
[24]
According to
Robert Rosen
in the 1960s, holistic biology had become passe by the early 20th century, as more empirical science dominated by molecular chemistry had become popular.
[18]
Echoing him forty years later in 2006 Kling writes that the success of
molecular biology
throughout the 20th century had suppressed holistic computational methods.
[14]
By 2011 the
National Institutes of Health
had made grant money available to support over ten systems biology centers in the United States,
[25]
but by 2012 Hunter writes that systems biology still has someway to go to achieve its full potential. Nonetheless, proponents hoped that it might once prove more useful in the future.
[26]
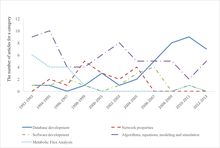 Shows trends in systems biology research by presenting the number of articles out of the top 30 cited systems biology papers during that time which include a specific topic
[27]
Shows trends in systems biology research by presenting the number of articles out of the top 30 cited systems biology papers during that time which include a specific topic
[27]
An important milestone in the development of systems biology has become the international project
Physiome
.
[
citation needed
]
Associated disciplines
[
edit
]
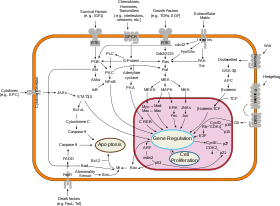 Overview of
signal transduction
pathways
Overview of
signal transduction
pathways
According to the interpretation of systems biology as using large data sets using interdisciplinary tools, a typical application is
metabolomics
, which is the complete set of all the metabolic products,
metabolites
, in the system at the organism, cell, or tissue level.
[28]
Items that may be a computer database include:
phenomics
, organismal variation in
phenotype
as it changes during its life span;
genomics
, organismal
deoxyribonucleic acid
(DNA) sequence, including intra-organismal cell specific variation. (i.e.,
telomere
length variation);
epigenomics
/
epigenetics
, organismal and corresponding cell specific transcriptomic regulating factors not empirically coded in the genomic sequence. (i.e.,
DNA methylation
,
Histone acetylation and deacetylation
, etc.);
transcriptomics
, organismal, tissue or whole cell
gene expression
measurements by
DNA microarrays
or
serial analysis of gene expression
;
interferomics
, organismal, tissue, or cell-level transcript correcting factors (i.e.,
RNA interference
),
proteomics
, organismal, tissue, or cell level measurements of proteins and peptides via
two-dimensional gel electrophoresis
,
mass spectrometry
or multi-dimensional protein identification techniques (advanced
HPLC
systems coupled with
mass spectrometry
). Sub disciplines include
phosphoproteomics
,
glycoproteomics
and other methods to detect chemically modified proteins;
glycomics
, organismal, tissue, or cell-level measurements of
carbohydrates
;
lipidomics
, organismal, tissue, or cell level measurements of
lipids
.
[
citation needed
]
The molecular interactions within the cell are also studied, this is called
interactomics
.
[29]
A discipline in this field of study is
protein?protein interactions
, although interactomics includes the interactions of other molecules.
[
citation needed
]
Neuroelectrodynamics
, where the computer's or a brain's computing function as a dynamic system is studied along with its (bio)physical mechanisms;
[30]
and
fluxomics
, measurements of the rates of metabolic reactions in a biological system (cell, tissue, or organism).
[28]
In approaching a systems biology problem there are two main approaches. These are the top down and bottom up approach. The top down approach takes as much of the system into account as possible and relies largely on experimental results. The
RNA-Seq
technique is an example of an experimental top down approach. Conversely, the bottom up approach is used to create detailed models while also incorporating experimental data. An example of the bottom up approach is the use of circuit models to describe a simple gene network.
[31]
Various technologies utilized to capture dynamic changes in mRNA, proteins, and post-translational modifications.
Mechanobiology
, forces and physical properties at all scales, their interplay with other regulatory mechanisms;
[32]
biosemiotics
, analysis of the system of
sign relations
of an organism or other biosystems;
Physiomics
, a systematic study of
physiome
in biology.
Cancer systems biology
is an example of the systems biology approach, which can be distinguished by the specific object of study (
tumorigenesis
and
treatment of cancer
). It works with the specific data (patient samples, high-throughput data with particular attention to characterizing
cancer genome
in patient tumour samples) and tools (immortalized cancer
cell lines
,
mouse models
of tumorigenesis,
xenograft
models,
high-throughput sequencing
methods, siRNA-based gene knocking down
high-throughput screenings
, computational modeling of the consequences of somatic
mutations
and
genome instability
).
[33]
The long-term objective of the systems biology of cancer is ability to better diagnose cancer, classify it and better predict the outcome of a suggested treatment, which is a basis for
personalized cancer medicine
and
virtual cancer patient
in more distant prospective. Significant efforts in computational systems biology of cancer have been made in creating realistic multi-scale
in silico
models of various tumours.
[34]
The systems biology approach often involves the development of
mechanistic
models, such as the reconstruction of
dynamic systems
from the quantitative properties of their elementary building blocks.
[35]
[36]
[37]
[38]
For instance, a cellular network can be modelled mathematically using methods coming from
chemical kinetics
[39]
and
control theory
. Due to the large number of parameters, variables and constraints in cellular networks, numerical and computational techniques are often used (e.g.,
flux balance analysis
).
[37]
[39]
Bioinformatics and data analysis
[
edit
]
Other aspects of computer science,
informatics
, and statistics are also used in systems biology. These include new forms of computational models, such as the use of
process calculi
to model biological processes (notable approaches include stochastic
π-calculus
, BioAmbients, Beta Binders, BioPEPA, and Brane calculus) and
constraint
-based modeling; integration of information from the literature, using techniques of
information extraction
and
text mining
;
[40]
development of online databases and repositories for sharing data and models, approaches to database integration and software interoperability via
loose coupling
of software, websites and databases, or commercial suits; network-based approaches for analyzing high dimensional genomic data sets. For example,
weighted correlation network analysis
is often used for identifying clusters (referred to as modules), modeling the relationship between clusters, calculating fuzzy measures of cluster (module) membership, identifying intramodular hubs, and for studying cluster preservation in other data sets; pathway-based methods for omics data analysis, e.g. approaches to identify and score pathways with differential activity of their gene, protein, or metabolite members.
[41]
Much of the analysis of genomic data sets also include identifying correlations. Additionally, as much of the information comes from different fields, the development of syntactically and semantically sound ways of representing biological models is needed.
[42]
Creating biological models
[
edit
]
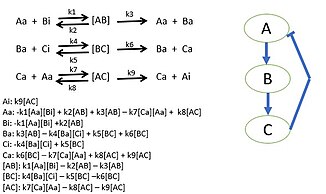 A simple three protein negative feedback loop modeled with mass action kinetic differential equations. Each protein interaction is described by a Michaelis?Menten reaction.
[43]
A simple three protein negative feedback loop modeled with mass action kinetic differential equations. Each protein interaction is described by a Michaelis?Menten reaction.
[43]
Researchers begin by choosing a biological pathway and diagramming all of the protein, gene, and/or metabolic pathways. After determining all of the interactions,
mass action kinetics
or
enzyme kinetic rate laws
are used to describe the speed of the reactions in the system. Using mass-conservation, the
differential equations
for the biological system can be constructed. Experiments or parameter fitting can be done to determine the parameter values to use in the
differential equations
.
[44]
These parameter values will be the various kinetic constants required to fully describe the model. This model determines the behavior of species in biological systems and bring new insight to the specific activities of system. Sometimes it is not possible to gather all reaction rates of a system. Unknown reaction rates are determined by simulating the model of known parameters and target behavior which provides possible parameter values.
[45]
[43]
The use of constraint-based reconstruction and analysis (COBRA) methods has become popular among systems biologists to simulate and predict the metabolic phenotypes, using genome-scale models. One of the methods is the
flux balance analysis
(FBA) approach, by which one can study the biochemical networks and analyze the flow of metabolites through a particular metabolic network, by optimizing the objective function of interest (e.g. maximizing biomass production to predict growth).
[46]
 Plot of Concentrations vs time for the simple three protein negative feedback loop. All parameters are set to either 0 or 1 for initial conditions. The reaction is allowed to proceed until it hits equilibrium. This plot is of the change in each protein over time.
Plot of Concentrations vs time for the simple three protein negative feedback loop. All parameters are set to either 0 or 1 for initial conditions. The reaction is allowed to proceed until it hits equilibrium. This plot is of the change in each protein over time.
See also
[
edit
]
References
[
edit
]
- ^
a
b
c
Tavassoly, Iman; Goldfarb, Joseph; Iyengar, Ravi (2018-10-04). "Systems biology primer: the basic methods and approaches".
Essays in Biochemistry
.
62
(4): 487?500.
doi
:
10.1042/EBC20180003
.
ISSN
0071-1365
.
PMID
30287586
.
S2CID
52922135
.
- ^
Zewail, Ahmed (2008).
Physical Biology: From Atoms to Medicine
. Imperial College Press. p. 339.
- ^
Longo, Giuseppe; Montevil, Mael (2014).
Perspectives on Organisms - Springer
. Lecture Notes in Morphogenesis.
doi
:
10.1007/978-3-642-35938-5
.
ISBN
978-3-642-35937-8
.
S2CID
27653540
.
- ^
Bu Z, Callaway DJ (2011). "Proteins MOVE! Protein dynamics and long-range allostery in cell signaling".
Protein Structure and Diseases
. Advances in Protein Chemistry and Structural Biology. Vol. 83. pp. 163?221.
doi
:
10.1016/B978-0-12-381262-9.00005-7
.
ISBN
978-0-123-81262-9
.
PMID
21570668
.
- ^
Snoep, Jacky L; Westerhoff, Hans V (2005). "From isolation to integration, a systems biology approach for building the Silicon Cell". In Alberghina, Lilia; Westerhoff, Hans V (eds.).
Systems Biology: Definitions and Perspectives
. Topics in Current Genetics. Vol. 13. Berlin: Springer-Verlag. pp. 13?30.
doi
:
10.1007/b106456
.
ISBN
978-3-540-22968-1
.
- ^
"Systems Biology: the 21st Century Science"
. Institute for Systems Biology
. Retrieved
15 June
2011
.
- ^
a
b
Noble, Denis
(2006).
The music of life: Biology beyond the genome
. Oxford: Oxford University Press. p. 176.
ISBN
978-0-19-929573-9
.
- ^
Sauer, Uwe; Heinemann, Matthias; Zamboni, Nicola (27 April 2007).
"Genetics: Getting Closer to the Whole Picture"
.
Science
.
316
(5824): 550?551.
doi
:
10.1126/science.1142502
.
PMID
17463274
.
S2CID
42448991
.
- ^
Kholodenko, Boris N; Sauro, Herbert M (2005). "Mechanistic and modular approaches to modeling and inference of cellular regulatory networks". In Alberghina, Lilia; Westerhoff, Hans V (eds.).
Systems Biology: Definitions and Perspectives
. Topics in Current Genetics. Vol. 13. Berlin: Springer-Verlag. pp. 357?451.
doi
:
10.1007/b136809
.
ISBN
978-3-540-22968-1
.
- ^
Chiara Romualdi; Gerolamo Lanfranchi (2009). "Statistical Tools for Gene Expression Analysis and Systems Biology and Related Web Resources". In Stephen Krawetz (ed.).
Bioinformatics for Systems Biology
(2nd ed.). Humana Press. pp. 181?205.
doi
:
10.1007/978-1-59745-440-7_11
.
ISBN
978-1-59745-440-7
.
- ^
Voit, Eberhard (2012).
A First Course in Systems Biology
. Garland Science.
ISBN
9780815344674
.
- ^
Baitaluk, M. (2009). "System Biology of Gene Regulation".
Biomedical Informatics
. Methods in Molecular Biology. Vol. 569. pp. 55?87.
doi
:
10.1007/978-1-59745-524-4_4
.
ISBN
978-1-934115-63-3
.
PMID
19623486
.
- ^
Wright, Sewall (1934).
"Physiological and Evolutionary Theories of Dominance"
.
The American Naturalist
. pp. 24?53.
- ^
a
b
c
Kling, Jim (3 March 2006).
"Working the Systems"
. Science
. Retrieved
15 June
2011
.
- ^
"HMS launches new department to study systems biology"
. Harvard Gazette. September 23, 2003.
- ^
Karr, Jonathan R.; Sanghvi, Jayodita C.; Macklin, Derek N.; Gutschow, Miriam V.; Jacobs, Jared M.; Bolival, Benjamin; Assad-Garcia, Nacyra; Glass, John I.; Covert, Markus W. (July 2012).
"A Whole-Cell Computational Model Predicts Phenotype from Genotype"
.
Cell
.
150
(2): 389?401.
doi
:
10.1016/j.cell.2012.05.044
.
PMC
3413483
.
PMID
22817898
.
- ^
Mesarovic, Mihajlo D.
(1968).
Systems Theory and Biology
. Berlin: Springer-Verlag.
- ^
a
b
Rosen, Robert
(5 July 1968). "A Means Toward a New Holism".
Science
.
161
(3836): 34?35.
Bibcode
:
1968Sci...161...34M
.
doi
:
10.1126/science.161.3836.34
.
JSTOR
1724368
.
- ^
Kacser, H; Burns, JA (1973). "The control of flux".
Symposia of the Society for Experimental Biology
.
27
: 65?104.
PMID
4148886
.
- ^
Kacser, H; Burns, JA; Fell, DA (1995). "The control of flux".
Biochemical Society Transactions
.
23
(2): 341?366.
doi
:
10.1042/bst0230341
.
PMID
7672373
.
- ^
Heinrich, R; Rapoport, TA (1974).
"A linear steady-state theory of enzymatic chains: general properties, control and effector strength"
.
European Journal of Biochemistry
.
42
(1): 89?95.
doi
:
10.1111/j.1432-1033.1974.tb03318.x
.
PMID
4830198
.
- ^
Savageau, Michael A. (December 1969).
"Journal of Theoretical Biology"
.
25
(3): 365?369.
doi
:
10.1016/S0022-5193(69)80026-3
.
PMID
5387046
.
- ^
Savageau, Michael A. (December 1969).
"Journal of Theoretical Biology"
.
25
(3): 370?379.
doi
:
10.1016/S0022-5193(69)80027-5
.
PMID
5387047
.
- ^
Savageau, Michael A. (February 1970).
"Journal of Theoretical Biology"
.
26
(2): 215?226.
doi
:
10.1016/S0022-5193(70)80013-3
.
PMID
5434343
.
- ^
"Systems Biology - National Institute of General Medical Sciences"
. Archived from
the original
on 19 October 2013
. Retrieved
12 December
2012
.
- ^
Hunter, Philip (May 2012).
"Back down to Earth: Even if it has not yet lived up to its promises, systems biology has now matured and is about to deliver its first results"
.
EMBO Reports
.
13
(5): 408?411.
doi
:
10.1038/embor.2012.49
.
PMC
3343359
.
PMID
22491028
.
- ^
Zou, Yawen; Laubichler, Manfred D. (2018-07-25).
"From systems to biology: A computational analysis of the research articles on systems biology from 1992 to 2013"
.
PLOS ONE
.
13
(7): e0200929.
Bibcode
:
2018PLoSO..1300929Z
.
doi
:
10.1371/journal.pone.0200929
.
ISSN
1932-6203
.
PMC
6059489
.
PMID
30044828
.
- ^
a
b
Cascante, Marta; Marin, Silvia (2008-09-30). "Metabolomics and fluxomics approaches".
Essays in Biochemistry
.
45
: 67?82.
doi
:
10.1042/bse0450067
.
ISSN
0071-1365
.
PMID
18793124
.
- ^
Cusick, Michael E.; Klitgord, Niels; Vidal, Marc; Hill, David E. (2005-10-15).
"Interactome: gateway into systems biology"
.
Human Molecular Genetics
.
14
(suppl_2): R171?R181.
doi
:
10.1093/hmg/ddi335
.
ISSN
0964-6906
.
PMID
16162640
.
- ^
Aur, Dorian (2012). "From Neuroelectrodynamics to Thinking Machines".
Cognitive Computation
.
4
(1): 4?12.
doi
:
10.1007/s12559-011-9106-3
.
ISSN
1866-9956
.
S2CID
12355069
.
- ^
Loor, Khuram Shahzad and Juan J. (2012-07-31).
"Application of Top-Down and Bottom-up Systems Approaches in Ruminant Physiology and Metabolism"
.
Current Genomics
.
13
(5): 379?394.
doi
:
10.2174/138920212801619269
.
PMC
3401895
.
PMID
23372424
.
- ^
Spill, Fabian; Bakal, Chris; Mak, Michael (2018).
"Mechanical and Systems Biology of Cancer"
.
Computational and Structural Biotechnology Journal
.
16
: 237?245.
arXiv
:
1807.08990
.
Bibcode
:
2018arXiv180708990S
.
doi
:
10.1016/j.csbj.2018.07.002
.
PMC
6077126
.
PMID
30105089
.
- ^
Barillot, Emmanuel; Calzone, Laurence; Hupe, Philippe; Vert, Jean-Philippe; Zinovyev, Andrei (2012).
Computational Systems Biology of Cancer
. Chapman & Hall/CRCMathematical & Computational Biology. p. 461.
ISBN
978-1439831441
.
- ^
Byrne, Helen M.
(2010). "Dissecting cancer through mathematics: from the cell to the animal model".
Nature Reviews Cancer
.
10
(3): 221?230.
doi
:
10.1038/nrc2808
.
PMID
20179714
.
S2CID
24616792
.
- ^
Gardner, Timothy .S; di Bernardo, Diego; Lorenz, David; Collins, James J. (4 July 2003). "Inferring Genetic Networks and Identifying Compound Mode of Action via Expression Profiling".
Science
.
301
(5629): 102?105.
Bibcode
:
2003Sci...301..102G
.
doi
:
10.1126/science.1081900
.
PMID
12843395
.
S2CID
8356492
.
- ^
di Bernardo, Diego; Thompson, Michael J.; Gardner, Timothy S.; Chobot, Sarah E.; Eastwood, Erin L.; Wojtovich, Andrew P.; Elliott, Sean J.; Schaus, Scott E.; Collins, James J. (March 2005). "Chemogenomic profiling on a genome-wide scale using reverse-engineered gene networks".
Nature Biotechnology
.
23
(3): 377?383.
doi
:
10.1038/nbt1075
.
PMID
15765094
.
S2CID
16270018
.
- ^
a
b
Tavassoly, Iman (2015).
Dynamics of Cell Fate Decision Mediated by the Interplay of Autophagy and Apoptosis in Cancer Cells
. Springer Theses. Springer International Publishing.
doi
:
10.1007/978-3-319-14962-2
.
ISBN
978-3-319-14961-5
.
S2CID
89307028
.
- ^
Korkut, A; Wang, W; Demir, E; Aksoy, BA; Jing, X; Molinelli, EJ; Babur, O; Bemis, DL; Onur Sumer, S; Solit, DB; Pratilas, CA; Sander, C (18 August 2015).
"Perturbation biology nominates upstream-downstream drug combinations in RAF inhibitor resistant melanoma cells"
.
eLife
.
4
.
doi
:
10.7554/eLife.04640
.
PMC
4539601
.
PMID
26284497
.
- ^
a
b
Gupta, Ankur; Rawlings, James B. (April 2014).
"Comparison of Parameter Estimation Methods in Stochastic Chemical Kinetic Models: Examples in Systems Biology"
.
AIChE Journal
.
60
(4): 1253?1268.
doi
:
10.1002/aic.14409
.
ISSN
0001-1541
.
PMC
4946376
.
PMID
27429455
.
- ^
Ananadou, Sophia
; Kell, Douglas; Tsujii, Jun-ichi (December 2006). "Text mining and its potential applications in systems biology".
Trends in Biotechnology
.
24
(12): 571?579.
doi
:
10.1016/j.tibtech.2006.10.002
.
PMID
17045684
.
- ^
Glaab, Enrico; Schneider, Reinhard (2012).
"PathVar: analysis of gene and protein expression variance in cellular pathways using microarray data"
.
Bioinformatics
.
28
(3): 446?447.
doi
:
10.1093/bioinformatics/btr656
.
PMC
3268235
.
PMID
22123829
.
- ^
Bardini, R.; Politano, G.; Benso, A.; Di Carlo, S. (2017-01-01).
"Multi-level and hybrid modelling approaches for systems biology"
.
Computational and Structural Biotechnology Journal
.
15
: 396?402.
doi
:
10.1016/j.csbj.2017.07.005
.
ISSN
2001-0370
.
PMC
5565741
.
PMID
28855977
.
- ^
a
b
Transtrum, Mark K.; Qiu, Peng (2016-05-17).
"Bridging Mechanistic and Phenomenological Models of Complex Biological Systems"
.
PLOS Computational Biology
.
12
(5): e1004915.
arXiv
:
1509.06278
.
Bibcode
:
2016PLSCB..12E4915T
.
doi
:
10.1371/journal.pcbi.1004915
.
ISSN
1553-7358
.
PMC
4871498
.
PMID
27187545
.
- ^
Chellaboina, V.; Bhat, S. P.; Haddad, W. M.; Bernstein, D. S. (August 2009). "Modeling and analysis of mass-action kinetics".
IEEE Control Systems Magazine
.
29
(4): 60?78.
doi
:
10.1109/MCS.2009.932926
.
ISSN
1941-000X
.
S2CID
12122032
.
- ^
Brown, Kevin S.; Sethna, James P. (2003-08-12). "Statistical mechanical approaches to models with many poorly known parameters".
Physical Review E
.
68
(2): 021904.
Bibcode
:
2003PhRvE..68b1904B
.
doi
:
10.1103/physreve.68.021904
.
ISSN
1063-651X
.
PMID
14525003
.
- ^
Orth, Jeffrey D; Thiele, Ines; Palsson, Bernhard Ø (March 2010).
"What is flux balance analysis?"
.
Nature Biotechnology
.
28
(3): 245?248.
doi
:
10.1038/nbt.1614
.
ISSN
1087-0156
.
PMC
3108565
.
PMID
20212490
.
Further reading
[
edit
]
- Klipp, Edda; Liebermeister, Wolfram; Wierling, Christoph; Kowald, Axel (2016).
Systems Biology - A Textbook, 2nd edition
. Wiley.
ISBN
978-3-527-33636-4
.
- Asfar S. Azmi, ed. (2012).
Systems Biology in Cancer Research and Drug Discovery
. Springer.
ISBN
978-94-007-4819-4
.
- Kitano, Hiroaki
(15 October 2001).
Foundations of Systems Biology
. MIT Press.
ISBN
978-0-262-11266-6
.
- Werner, Eric (29 March 2007).
"All systems go"
.
Nature
.
446
(7135): 493?494.
Bibcode
:
2007Natur.446..493W
.
doi
:
10.1038/446493a
.
provides a comparative review of three books:
- Alon, Uri
(7 July 2006).
An Introduction to Systems Biology: Design Principles of Biological Circuits
. Chapman & Hall.
ISBN
978-1-58488-642-6
.
- Kaneko, Kunihiko (15 September 2006).
Life: An Introduction to Complex Systems Biology
. Springer-Verlag.
Bibcode
:
2006lics.book.....K
.
ISBN
978-3-540-32666-3
.
- Palsson, Bernhard O. (16 January 2006).
Systems Biology: Properties of Reconstructed Networks
. Cambridge University Press.
ISBN
978-0-521-85903-5
.
- Werner Dubitzky; Olaf Wolkenhauer; Hiroki Yokota; Kwan-Hyun Cho, eds. (13 August 2013).
Encyclopedia of Systems Biology
. Springer-Verlag.
ISBN
978-1-4419-9864-4
.
External links
[
edit
]