Species of coronavirus causing SARS and COVID-19
This article is about a species of coronavirus comprising multiple strains. For the strain that causes SARS, see
SARS-CoV-1
. For the strain that causes COVID-19, see
SARS-CoV-2
.
Betacoronavirus pandemicum
[2]
(also known as
Severe acute respiratory syndrome?related coronavirus
, abbreviated as
SARSr-CoV
or
SARS-CoV
)
[note 1]
is a species of virus consisting of many known strains. Two
strains
of the virus have caused
outbreaks
of severe
respiratory diseases
in humans:
severe acute respiratory syndrome coronavirus 1
(SARS-CoV or SARS-CoV-1), the cause of
the 2002?2004 outbreak
of
severe acute respiratory syndrome
(SARS), and
severe acute respiratory syndrome coronavirus 2
(SARS-CoV-2), the cause of
the pandemic
of
COVID-19
.
[3]
[4]
There are hundreds of other strains of SARSr-CoV, which are only known to infect non-human mammal species:
bats are a major reservoir
of many strains of SARSr-CoV; several strains have been identified in
Himalayan palm civets
, which were likely ancestors of SARS-CoV-1.
[3]
[5]
[6]
[7]
These
enveloped
,
positive-sense single-stranded RNA viruses
enter host cells by binding to the
angiotensin-converting enzyme 2
(ACE2) receptor.
[8]
The SARSr-CoV species is a member of the genus
Betacoronavirus
and the only species of the subgenus
Sarbecovirus
(
SAR
S
Be
ta
co
rona
virus
).
[9]
[10]
The SARS-related coronavirus was one of several viruses identified by the
World Health Organization
(WHO) in 2016 as a likely cause of a future
epidemic
in a new plan developed after the
Ebola epidemic
for urgent research and development before and during an epidemic towards
diagnostic tests
,
vaccines
and
medicines
. This prediction came to pass with the
COVID-19 pandemic
.
[11]
[12]
Classification
[
edit
]
SARS-related coronavirus is a member of the genus
Betacoronavirus
(group 2) and
monotypic
of the subgenus
Sarbecovirus
(subgroup B).
[13]
Sarbecoviruses, unlike
embecoviruses
or
alphacoronaviruses
, have only one
papain-like proteinase
(PLpro) instead of two in the
open reading frame
ORF1ab
.
[14]
SARSr-CoV was determined to be an early split-off from the betacoronaviruses based on a set of conserved domains that it shares with the group.
[15]
[16]
Bats serve as the main host reservoir species for the SARS-related coronaviruses like SARS-CoV-1 and SARS-CoV-2. The virus has coevolved in the
bat host reservoir
over a long period of time.
[17]
Only recently have strains of SARS-related coronavirus been observed to have evolved into having been able to make the
cross-species jump
from bats to humans, as in the case of the strains
SARS-CoV-1
and
SARS-CoV-2
.
[18]
[8]
Both of these strains descended from a single ancestor but made the cross-species jump into humans separately. SARS-CoV-2 is not a direct descendant of SARS-CoV-1.
[3]
Genome
[
edit
]
 Genome
organization and viral
proteins
of
SARS-CoV
Genome
organization and viral
proteins
of
SARS-CoV
The SARS-related
coronavirus
is an enveloped,
positive-sense, single-stranded RNA virus
. Its genome is about 30
kb
, which is one of the largest among RNA viruses. The virus has 14
open reading frames
which overlap in some cases.
[19]
The genome has the usual
5′ methylated cap
and a
3′ polyadenylated tail
.
[20]
There are 265
nucleotides
in the
5'UTR
and 342 nucleotides in the
3'UTR
.
[19]
The 5' methylated cap and 3' polyadenylated tail allows the
positive-sense RNA genome
to be directly
translated
by the host cell's
ribosome
on
viral entry
.
[21]
SARSr-CoV is similar to other coronaviruses in that its genome expression starts with translation by the host cell's ribosomes of its initial two large overlapping open reading frames (ORFs), 1a and 1b, both of which produce
polyproteins
.
[19]
Function of SARSr-CoV
genome proteins
|
| Protein
|
Function
[22]
[23]
[24]
[25]
|
ORF1ab
P0C6X7
|
Replicase/transcriptase polyprotein (pp1ab)
(
nonstructural proteins
)
|
ORF2
P59594
|
Spike
(S) protein, virus binding and entry
(
structural protein
)
|
ORF3a
P59632
|
Interacts with S, E, M structural proteins;
Ion channel
activity;
Upregulates
cytokines
and
chemokines
such as
IL-8
and
RANTES
;
Upregulates
NF-κB
and
JNK
;
Induces
apoptosis
and
cell cycle arrest
, via
Caspase 8
and
-9
,
and by
Bax
,
p53
, and
p38 MAP kinase
|
ORF3b
P59633
|
Upregulates
cytokines
and
chemokines
by
RUNX1b
;
Inhibits
Type I IFN
production and signaling;
Induces apoptosis and
cell cycle arrest
;
|
ORF3c
P0DTG1
|
Unknown; first identified in SARS-CoV-2 but also present in SARS-CoV
|
ORF3d
P0DTG0
|
Novel gene in SARS-CoV-2, of unknown function
|
ORF4
P59637
|
Envelope
(E) protein, virus assembly and budding
(
structural protein
)
|
ORF5
P59596
|
Membrane
(M) protein, virus assembly and budding
(
structural protein
)
|
ORF6
P59634
|
Enhances cellular DNA synthesis;
Inhibits Type I IFN production and signaling
|
ORF7a
P59635
|
Inhibits cellular protein synthesis;
Induces inflammatory response by
NF-kappaB
and
IL-8 promotor
;
Upregulate chemokines such as IL-8 and RANTES;
Upregulates JNK, p38 MAP kinase;
Induces apoptosis and cell cycle arrest
|
ORF7b
Q7TFA1
|
Unknown
|
ORF8a
Q7TFA0
|
Induces apoptosis through
mitochondria
pathway
|
ORF8b
Q80H93
|
Enhances cellular DNA synthesis, also known as X5.
|
ORF9a
P59595
|
Nucleocapsid
(N) protein, viral RNA packaging
(
structural protein
)
|
ORF9b
P59636
|
Induces apoptosis
|
ORF9c
Q7TLC7
|
Also known as ORF14; function unknown and may not be protein-coding
|
ORF10
A0A663DJA2
|
Novel gene in SARS-CoV-2, of unknown function; may not be protein-coding
|
| UniProt
identifiers shown for
SARS-CoV
proteins unless they are specific to SARS-CoV-2
|
The functions of several of the viral proteins are known.
[26]
ORFs 1a and 1b encode the replicase/transcriptase polyprotein, and later ORFs 2, 4, 5, and 9a encode, respectively, the four major structural proteins:
spike
(S),
envelope
(E),
membrane
(M), and
nucleocapsid
(N).
[27]
The later ORFs also encode for eight unique proteins (orf3a to orf9b), known as the
accessory proteins
, many with no known homologues. The different functions of the accessory proteins are not well understood.
[26]
SARS coronaviruses have been genetically engineered in several laboratories.
[28]
Phylogenetics
[
edit
]
 Phylogenetic tree
of SARS-CoV-2 and closely related betacoronaviruses (left) and their geographic context (right)
Phylogenetic tree
of SARS-CoV-2 and closely related betacoronaviruses (left) and their geographic context (right)
Phylogenetic analysis
showed that the evolutionary branch composed of Bat coronavirus BtKY72 and BM48-31 was the base group of SARS?related CoVs evolutionary tree, which separated from other SARS?related CoVs earlier than SARS-CoV-1 and SARS-CoV-2.
[29]
[3]
| SARSr‑CoV
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
SARS-CoV-1 related coronavirus
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
SARS-CoV-2 related coronavirus
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
SARS-CoV-1 related
[
edit
]
A phylogenetic tree based on whole-genome sequences of SARS-CoV-1 and related coronaviruses is:
SARS-CoV-2 related
[
edit
]
A phylogenetic tree based on whole-genome sequences of SARS-CoV-2 and related coronaviruses is:
[39]
[40]
| SARS‑CoV‑2 related coronavirus
|
|
|
(
Bat
)
Rc-o319
, 81% to SARS-CoV-2,
Rhinolophus cornutus
,
Iwate
, Japan
[41]
|
|
|
|
|
|
|
|
|
|
|
|
|
|
Pangolin
SARSr-CoV-GX, 85.3% to SARS-CoV-2,
Manis javanica
, smuggled from Southeast Asia
[43]
|
|
|
|
|
|
|
Pangolin SARSr-CoV-GD, 90.1% to SARS-CoV-2,
Manis javanica
, smuggled from Southeast Asia
[44]
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
SARS-CoV-1
, 79% to SARS-CoV-2
|
|
|
Morphology
[
edit
]
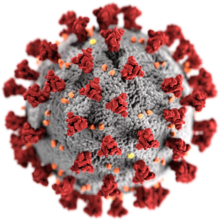 Illustration created at the
Centers for Disease Control and Prevention
(CDC), reveals ultrastructural morphology exhibited by coronaviruses; note the
spikes
that adorn the outer surface, which impart the look of a corona surrounding the
virion
.
[49]
Illustration created at the
Centers for Disease Control and Prevention
(CDC), reveals ultrastructural morphology exhibited by coronaviruses; note the
spikes
that adorn the outer surface, which impart the look of a corona surrounding the
virion
.
[49]
 Illustration of SARSr-CoV
virion
Illustration of SARSr-CoV
virion
The morphology of the SARS-related coronavirus is characteristic of the coronavirus family as a whole. The viruses are large
pleomorphic
spherical particles with bulbous surface projections that form a corona around the particles in electron micrographs.
[50]
The size of the virus particles is in the 80?90 nm range. The envelope of the virus in electron micrographs appears as a distinct pair of electron dense shells.
[51]
The
viral envelope
consists of a
lipid bilayer
where the
membrane
(M),
envelope
(E) and
spike
(S) proteins are anchored.
[52]
The spike proteins provide the virus with its bulbous surface projections, known as
peplomers
. The spike protein's interaction with its complement
host cell receptor
is central in determining the
tissue tropism
,
infectivity
, and
species range
of the virus.
[53]
[54]
Inside the envelope, there is the
nucleocapsid
, which is formed from multiple copies of the
nucleocapsid
(N) protein, which are bound to the positive-sense single-stranded (~30
kb
) RNA genome in a continuous
beads-on-a-string
type conformation.
[55]
[56]
The lipid bilayer envelope, membrane proteins, and nucleocapsid protect the virus when it is outside the host.
[57]
Life cycle
[
edit
]
SARS-related coronavirus follows the replication strategy typical of all coronaviruses.
[20]
[58]
Attachment and entry
[
edit
]
 Coronavirus
replication
cycle
Coronavirus
replication
cycle
The attachment of the SARS-related coronavirus to the host cell is mediated by the spike protein and its receptor.
[59]
The spike protein receptor binding domain (RBD) recognizes and attaches to the
angiotensin-converting enzyme 2
(ACE2) receptor.
[8]
Following attachment, the virus can enter the host cell by two different paths. The path the virus takes depends on the host
protease
available to cleave and activate the receptor-attached spike protein.
[60]
The attachment of sarbecoviruses to ACE2 has been shown to be an
evolutionarily conserved
feature, present in many species of the taxon.
[61]
The first path the SARS coronavirus can take to enter the host cell is by
endocytosis
and uptake of the virus in an
endosome
. The receptor-attached spike protein is then activated by the host's pH-dependent
cysteine protease
cathepsin L
. Activation of the receptor-attached spike protein causes a
conformational change
, and the subsequent fusion of the viral envelope with the
endosomal wall
.
[60]
Alternatively, the virus can enter the host cell directly by
proteolytic
cleavage of the receptor-attached spike protein by the host's
TMPRSS2
or
TMPRSS11D
serine proteases
at the cell surface.
[62]
[63]
In the SARS coronavirus, the activation of the
C-terminal part
of the spike protein triggers the fusion of the viral envelope with the host cell membrane by inducing conformational changes which are not fully understood.
[64]
Genome translation
[
edit
]
After fusion the nucleocapsid passes into the
cytoplasm
, where the viral genome is released.
[59]
The genome
acts as a messenger RNA
, and the cell's ribosome
translates
two-thirds of the genome, which corresponds to the open reading frame
ORF1a
and
ORF1b
, into two large overlapping polyproteins, pp1a and pp1ab.
The larger polyprotein pp1ab is a result of a
-1 ribosomal frameshift
caused by a
slippery sequence
(UUUAAAC) and a downstream
RNA pseudoknot
at the end of open reading frame ORF1a.
[67]
The ribosomal frameshift allows for the continuous translation of ORF1a followed by ORF1b.
[68]
The polyproteins contain their own
proteases
,
PLpro
and
3CLpro
, which cleave the polyproteins at different specific sites. The cleavage of polyprotein pp1ab yields 16 nonstructural proteins (nsp1 to nsp16). Product proteins include various replication proteins such as
RNA-dependent RNA polymerase
(RdRp),
RNA helicase
, and
exoribonuclease
(ExoN).
[68]
The two SARS-CoV-2 proteases (PLpro and 3CLpro) also interfere with the immune system response to the viral infection by cleaving three immune system proteins. PLpro cleaves
IRF3
and 3CLpro cleaves both
NLRP12
and
TAB1
. "Direct cleavage of IRF3 by NSP3 could explain the blunted Type-I IFN response seen during SARS-CoV-2 infections while NSP5 mediated cleavage of NLRP12 and TAB1 point to a molecular mechanism for enhanced production of IL-6 and inflammatory response observed in COVID-19 patients."
[69]
Replication and transcription
[
edit
]
 Model of the
replicase
-transcriptase complex of a
coronavirus
.
RdRp
for
replication
(red),
ExoN
for
proofreading
(dark blue), ExoN cofactor (yellow),
RBPs
to avoid
secondary structure
(light blue),
RNA sliding clamp
for
processivity
and
primase
domain
for
priming
(green/orange), and a
helicase
to unwind RNA (downstream).
Model of the
replicase
-transcriptase complex of a
coronavirus
.
RdRp
for
replication
(red),
ExoN
for
proofreading
(dark blue), ExoN cofactor (yellow),
RBPs
to avoid
secondary structure
(light blue),
RNA sliding clamp
for
processivity
and
primase
domain
for
priming
(green/orange), and a
helicase
to unwind RNA (downstream).
A number of the nonstructural replication proteins coalesce to form a
multi-protein
replicase-transcriptase complex (RTC).
[68]
The main replicase-transcriptase protein is the
RNA-dependent RNA polymerase
(RdRp). It is directly involved in the
replication
and
transcription
of RNA from an RNA strand. The other nonstructural proteins in the complex assist in the replication and transcription process.
[65]
The protein nsp14 is a
3'-5' exoribonuclease
which provides extra fidelity to the replication process. The exoribonuclease provides a
proofreading
function to the complex which the RNA-dependent RNA polymerase lacks. Similarly, proteins nsp7 and nsp8 form a hexadecameric sliding clamp as part of the complex which greatly increases the
processivity
of the RNA-dependent RNA polymerase.
[65]
The coronaviruses require the increased fidelity and processivity during RNA synthesis because of the relatively large genome size in comparison to other RNA viruses.
[70]
One of the main functions of the replicase-transcriptase complex is to transcribe the viral genome. RdRp directly mediates the
synthesis
of negative-sense
subgenomic RNA
molecules from the positive-sense genomic RNA. This is followed by the transcription of these negative-sense subgenomic RNA molecules to their corresponding positive-sense
mRNAs
.
[71]
The other important function of the replicase-transcriptase complex is to replicate the viral genome. RdRp directly mediates the
synthesis
of negative-sense genomic RNA from the positive-sense genomic RNA. This is followed by the replication of positive-sense genomic RNA from the negative-sense genomic RNA.
[71]
The replicated positive-sense genomic RNA becomes the genome of the
progeny viruses
. The various smaller mRNAs are transcripts from the last third of the virus genome which follows the reading frames ORF1a and ORF1b. These mRNAs are translated into the four structural proteins (S, E, M, and N) that will become part of the progeny virus particles and also eight other accessory proteins (orf3 to orf9b) which assist the virus.
[72]
Recombination
[
edit
]
When two SARS-CoV
genomes
are present in a host cell, they may interact with each other to form recombinant genomes that can be transmitted to progeny viruses. Recombination likely occurs during genome replication when the
RNA polymerase
switches from one template to another (copy choice recombination).
[73]
Human SARS-CoV appears to have had a complex history of
recombination
between ancestral
coronaviruses
that were hosted in several different animal groups.
[73]
[74]
Assembly and release
[
edit
]
RNA translation
occurs inside the
endoplasmic reticulum
. The viral structural proteins S, E and M move along the secretory pathway into the
Golgi intermediate compartment
. There, the M proteins direct most protein-protein interactions required for assembly of viruses following its binding to the nucleocapsid.
[75]
Progeny viruses are released from the host cell by
exocytosis
through secretory vesicles.
[75]
See also
[
edit
]
Notes
[
edit
]
- ^
The terms
SARSr-CoV
and
SARS-CoV
are sometimes used interchangeably, especially prior to the discovery of SARS-CoV-2. This may cause confusion when some publications refer to SARS-CoV-1 as
SARS-CoV
.
References
[
edit
]
- ^
"ICTV Taxonomy history:
Severe acute respiratory syndrome-related coronavirus
"
.
International Committee on Taxonomy of Viruses (ICTV)
. Retrieved
27 January
2019
.
- ^
"Taxon Details | ICTV"
.
International Committee on Taxonomy of Viruses (ICTV)
. Retrieved
30 May
2024
.
- ^
a
b
c
d
Coronaviridae Study Group of the International Committee on Taxonomy of Viruses (March 2020).
"The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2"
.
Nature Microbiology
.
5
(4): 536?544.
doi
:
10.1038/s41564-020-0695-z
.
PMC
7095448
.
PMID
32123347
.
- ^
Kohen, Jon; Kupferschmidth, Kai (28 February 2020). "Strategies shift as coronavirus pandemic looms".
Science
.
367
(6481): 962?963.
Bibcode
:
2020Sci...367..962C
.
doi
:
10.1126/science.367.6481.962
.
PMID
32108093
.
S2CID
211556915
.
- ^
Lau SK, Li KS, Huang Y, Shek CT, Tse H, Wang M, et al. (March 2010).
"Ecoepidemiology and complete genome comparison of different strains of severe acute respiratory syndrome-related Rhinolophus bat coronavirus in China reveal bats as a reservoir for acute, self-limiting infection that allows recombination events"
.
Journal of Virology
.
84
(6): 2808?19.
doi
:
10.1128/JVI.02219-09
.
PMC
2826035
.
PMID
20071579
.
- ^
Branswell H (9 November 2015).
"SARS-like virus in bats shows potential to infect humans, study finds"
.
Stat News
. Retrieved
20 February
2020
.
- ^
Wong AC, Li X, Lau SK, Woo PC (February 2019).
"Global Epidemiology of Bat Coronaviruses"
.
Viruses
.
11
(2): 174.
doi
:
10.3390/v11020174
.
PMC
6409556
.
PMID
30791586
.
Most notably, horseshoe bats were found to be the reservoir of SARS-like CoVs, while palm civet cats are considered to be the intermediate host for SARS-CoVs [43,44,45].
- ^
a
b
c
Ge XY, Li JL, Yang XL, Chmura AA, Zhu G, Epstein JH, et al. (November 2013).
"Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor"
.
Nature
.
503
(7477): 535?8.
Bibcode
:
2013Natur.503..535G
.
doi
:
10.1038/nature12711
.
PMC
5389864
.
PMID
24172901
.
- ^
"Virus Taxonomy: 2018 Release"
.
International Committee on Taxonomy of Viruses (ICTV)
. October 2018
. Retrieved
13 January
2019
.
- ^
Woo PC, Huang Y, Lau SK, Yuen KY (August 2010).
"Coronavirus genomics and bioinformatics analysis"
.
Viruses
.
2
(8): 1804?20.
doi
:
10.3390/v2081803
.
PMC
3185738
.
PMID
21994708
.
Figure 2. Phylogenetic analysis of RNA-dependent RNA polymerases (Pol) of coronaviruses with complete genome sequences available. The tree was constructed by the neighbor-joining method and rooted using Breda virus polyprotein.
- ^
Kieny MP.
"After Ebola, a Blueprint Emerges to Jump-Start R&D"
.
Scientific American Blog Network
.
Archived
from the original on 20 December 2016
. Retrieved
13 December
2016
.
- ^
"LIST OF PATHOGENS"
.
World Health Organization
. Archived from
the original
on 20 December 2016
. Retrieved
13 December
2016
.
- ^
Wong AC, Li X, Lau SK, Woo PC (February 2019).
"Global Epidemiology of Bat Coronaviruses"
.
Viruses
.
11
(2): 174.
doi
:
10.3390/v11020174
.
PMC
6409556
.
PMID
30791586
.
See Figure 1.
- ^
Woo PC, Huang Y, Lau SK, Yuen KY (August 2010).
"Coronavirus genomics and bioinformatics analysis"
.
Viruses
.
2
(8): 1804?20.
doi
:
10.3390/v2081803
.
PMC
3185738
.
PMID
21994708
.
See Figure 1.
- ^
Woo PC, Huang Y, Lau SK, Yuen KY (August 2010).
"Coronavirus genomics and bioinformatics analysis"
.
Viruses
.
2
(8): 1804?20.
doi
:
10.3390/v2081803
.
PMC
3185738
.
PMID
21994708
.
Furthermore, subsequent phylogenetic analysis using both complete genome sequence and proteomic approaches, it was concluded that SARSr-CoV is probably an early split-off from the Betacoronavirus lineage [1]; See Figure 2.
- ^
"Coronaviridae - Figures - Positive Sense RNA Viruses - Positive Sense RNA Viruses (2011)"
.
International Committee on Taxonomy of Viruses (ICTV)
. Archived from
the original
on 3 April 2020
. Retrieved
6 March
2020
.
See Figure 2.
- ^
Gouilh MA, Puechmaille SJ, Gonzalez JP, Teeling E, Kittayapong P, Manuguerra JC (October 2011).
"SARS-Coronavirus ancestor's foot-prints in South-East Asian bat colonies and the refuge theory"
.
Infection, Genetics and Evolution
.
11
(7): 1690?702.
doi
:
10.1016/j.meegid.2011.06.021
.
PMC
7106191
.
PMID
21763784
.
Betacoronaviruses-b ancestors, meaning SARSr-CoVs ancestors, could have been historically hosted by the common ancestor of the Rhinolophidae and Hipposideridae and could have later evolved independently in the lineages leading towards Rhinolophidae and Hipposideridae betacoronaviruses.
- ^
Cui J, Han N, Streicker D, Li G, Tang X, Shi Z, et al. (October 2007).
"Evolutionary relationships between bat coronaviruses and their hosts"
.
Emerging Infectious Diseases
.
13
(10): 1526?32.
doi
:
10.3201/eid1310.070448
.
PMC
2851503
.
PMID
18258002
.
- ^
a
b
c
Snijder EJ, Bredenbeek PJ, Dobbe JC, Thiel V, Ziebuhr J, Poon LL, et al. (August 2003).
"Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage"
.
Journal of Molecular Biology
.
331
(5): 991?1004.
doi
:
10.1016/S0022-2836(03)00865-9
.
PMC
7159028
.
PMID
12927536
.
The SARS-CoV genome is ~29.7 kb long and contains 14 open reading frames (ORFs) flanked by 5′ and 3′-untranslated regions of 265 and 342 nucleotides, respectively (Figure 1).
- ^
a
b
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
- ^
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
- ^
McBride R, Fielding BC (November 2012).
"The role of severe acute respiratory syndrome (SARS)-coronavirus accessory proteins in virus pathogenesis"
.
Viruses
.
4
(11): 2902?23.
doi
:
10.3390/v4112902
.
PMC
3509677
.
PMID
23202509
.
See Table 1.
- ^
Tang X, Li G, Vasilakis N, Zhang Y, Shi Z, Zhong Y, Wang LF, Zhang S (March 2009).
"Differential stepwise evolution of SARS coronavirus functional proteins in different host species"
.
BMC Evolutionary Biology
.
9
(1): 52.
Bibcode
:
2009BMCEE...9...52T
.
doi
:
10.1186/1471-2148-9-52
.
PMC
2676248
.
PMID
19261195
.
- ^
Narayanan, Krishna; Huang, Cheng; Makino, Shinji (April 2008).
"SARS coronavirus Accessory Proteins"
.
Virus Research
.
133
(1): 113?121.
doi
:
10.1016/j.virusres.2007.10.009
.
ISSN
0168-1702
.
PMC
2720074
.
PMID
18045721
.
See Table 1.
- ^
Redondo, Natalia; Zaldivar-Lopez, Sara; Garrido, Juan J.; Montoya, Maria (7 July 2021).
"SARS-CoV-2 Accessory Proteins in Viral Pathogenesis: Knowns and Unknowns"
.
Frontiers in Immunology
.
12
: 708264.
doi
:
10.3389/fimmu.2021.708264
.
PMC
8293742
.
PMID
34305949
.
- ^
a
b
McBride R, Fielding BC (November 2012).
"The role of severe acute respiratory syndrome (SARS)-coronavirus accessory proteins in virus pathogenesis"
.
Viruses
.
4
(11): 2902?23.
doi
:
10.3390/v4112902
.
PMC
3509677
.
PMID
23202509
.
- ^
Snijder EJ, Bredenbeek PJ, Dobbe JC, Thiel V, Ziebuhr J, Poon LL, et al. (August 2003).
"Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage"
.
Journal of Molecular Biology
.
331
(5): 991?1004.
doi
:
10.1016/S0022-2836(03)00865-9
.
PMC
7159028
.
PMID
12927536
.
See Figure 1.
- ^
Kaina, Bernd (2021).
"On the Origin of SARS-CoV-2: Did Cell Culture Experiments Lead to Increased Virulence of the Progenitor Virus for Humans?"
.
In Vivo
.
35
(3): 1313?1326.
doi
:
10.21873/invivo.12384
.
PMC
8193286
.
PMID
33910809
.
- ^
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H; et al. (2020).
"Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding"
.
Lancet
.
395
(10224): 565?574.
doi
:
10.1016/S0140-6736(20)30251-8
.
PMC
7159086
.
PMID
32007145
.
{{
cite journal
}}
: CS1 maint: multiple names: authors list (
link
)
- ^
Kim, Yongkwan; Son, Kidong; Kim, Young-Sik; Lee, Sook-Young; Jheong, Weonhwa; Oem, Jae-Ku (2019).
"Complete genome analysis of a SARS-like bat coronavirus identified in the Republic of Korea"
.
Virus Genes
.
55
(4): 545?549.
doi
:
10.1007/s11262-019-01668-w
.
PMC
7089380
.
PMID
31076983
.
- ^
Xu, L; Zhang, F; Yang, W; Jiang, T; Lu, G; He, B; Li, X; Hu, T; Chen, G; Feng, Y; Zhang, Y; Fan, Q; Feng, J; Zhang, H; Tu, C (February 2016).
"Detection and characterization of diverse alpha- and betacoronaviruses from bats in China"
.
Virologica Sinica
.
31
(1): 69?77.
doi
:
10.1007/s12250-016-3727-3
.
PMC
7090707
.
PMID
26847648
.
- ^
a
b
Li, W. (2005).
"Bats Are Natural Reservoirs of SARS-Like Coronaviruses"
.
Science
.
310
(5748): 676?679.
Bibcode
:
2005Sci...310..676L
.
doi
:
10.1126/science.1118391
.
ISSN
0036-8075
.
PMID
16195424
.
S2CID
2971923
.
- ^
a
b
Xing?Yi Ge; Ben Hu; Zheng?Li Shi (2015). "BAT CORONAVIRUSES". In Lin-Fa Wang; Christopher Cowled (eds.).
Bats and Viruses: A New Frontier of Emerging Infectious Diseases
(First ed.). John Wiley & Sons. pp. 127?155.
doi
:
10.1002/9781118818824.ch5
.
- ^
He, Biao; Zhang, Yuzhen; Xu, Lin; Yang, Weihong; Yang, Fanli; Feng, Yun; et al. (2014).
"Identification of diverse alphacoronaviruses and genomic characterization of a novel severe acute respiratory syndrome-like coronavirus from bats in China"
.
J Virol
.
88
(12): 7070?82.
doi
:
10.1128/JVI.00631-14
.
PMC
4054348
.
PMID
24719429
.
- ^
a
b
Lau, Susanna K. P.; Feng, Yun; Chen, Honglin; Luk, Hayes K. H.; Yang, Wei-Hong; Li, Kenneth S. M.; Zhang, Yu-Zhen; Huang, Yi; et al. (2015).
"Severe Acute Respiratory Syndrome (SARS) Coronavirus ORF8 Protein Is Acquired from SARS-Related Coronavirus from Greater Horseshoe Bats through Recombination"
.
Journal of Virology
.
89
(20): 10532?10547.
doi
:
10.1128/JVI.01048-15
.
ISSN
0022-538X
.
PMC
4580176
.
PMID
26269185
.
- ^
a
b
Xing-Yi Ge; Jia-Lu Li; Xing-Lou Yang; et al. (2013).
"Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor"
.
Nature
.
503
(7477): 535?8.
Bibcode
:
2013Natur.503..535G
.
doi
:
10.1038/nature12711
.
PMC
5389864
.
PMID
24172901
.
- ^
Yang, Xing-Lou; Hu, Ben; Wang, Bo; Wang, Mei-Niang; Zhang, Qian; Zhang, Wei; et al. (2016).
"Isolation and Characterization of a Novel Bat Coronavirus Closely Related to the Direct Progenitor of Severe Acute Respiratory Syndrome Coronavirus"
.
Journal of Virology
.
90
(6): 3253?6.
doi
:
10.1128/JVI.02582-15
.
PMC
4810638
.
PMID
26719272
.
- ^
Ben, Hu; Hua, Guo; Peng, Zhou; Zheng-Li, Shi (2020).
"Characteristics of SARS-CoV-2 and COVID-19"
.
Nature Reviews Microbiology
.
19
(3): 141?154.
doi
:
10.1038/s41579-020-00459-7
.
PMC
7537588
.
PMID
33024307
.
- ^
a
b
Zhou H, Ji J, Chen X, Bi Y, Li J, Wang Q, et al. (August 2021).
"Identification of novel bat coronaviruses sheds light on the evolutionary origins of SARS-CoV-2 and related viruses"
.
Cell
.
184
(17): 4380?4391.e14.
doi
:
10.1016/j.cell.2021.06.008
.
PMC
8188299
.
PMID
34147139
.
- ^
a
b
Wacharapluesadee S, Tan CW, Maneeorn P, Duengkae P, Zhu F, Joyjinda Y, et al. (February 2021).
"Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia"
.
Nature Communications
.
12
(1): 972.
Bibcode
:
2021NatCo..12..972W
.
doi
:
10.1038/s41467-021-21240-1
.
PMC
7873279
.
PMID
33563978
.
- ^
Murakami S, Kitamura T, Suzuki J, Sato R, Aoi T, Fujii M, et al. (December 2020).
"Detection and Characterization of Bat Sarbecovirus Phylogenetically Related to SARS-CoV-2, Japan"
.
Emerging Infectious Diseases
.
26
(12): 3025?3029.
doi
:
10.3201/eid2612.203386
.
PMC
7706965
.
PMID
33219796
.
- ^
a
b
Zhou H, Chen X, Hu T, Li J, Song H, Liu Y, et al. (June 2020).
"A Novel Bat Coronavirus Closely Related to SARS-CoV-2 Contains Natural Insertions at the S1/S2 Cleavage Site of the Spike Protein"
.
Current Biology
.
30
(11): 2196?2203.e3.
doi
:
10.1016/j.cub.2020.05.023
.
PMC
7211627
.
PMID
32416074
.
- ^
Lam TT, Jia N, Zhang YW, Shum MH, Jiang JF, Zhu HC, et al. (July 2020). "Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins".
Nature
.
583
(7815): 282?285.
Bibcode
:
2020Natur.583..282L
.
doi
:
10.1038/s41586-020-2169-0
.
PMID
32218527
.
S2CID
214683303
.
- ^
Xiao K, Zhai J, Feng Y, Zhou N, Zhang X, Zou JJ, et al. (July 2020). "Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins".
Nature
.
583
(7815): 286?289.
Bibcode
:
2020Natur.583..286X
.
doi
:
10.1038/s41586-020-2313-x
.
PMID
32380510
.
S2CID
256822274
.
- ^
a
b
Delaune D, Hul V, Karlsson EA, Hassanin A, Ou TP, Baidaliuk A, et al. (November 2021).
"A novel SARS-CoV-2 related coronavirus in bats from Cambodia"
.
Nature Communications
.
12
(1): 6563.
Bibcode
:
2021NatCo..12.6563D
.
doi
:
10.1038/s41467-021-26809-4
.
PMC
8578604
.
PMID
34753934
.
- ^
Zhou H, Chen X, Hu T, Li J, Song H, Liu Y, et al. (June 2020).
"A Novel Bat Coronavirus Closely Related to SARS-CoV-2 Contains Natural Insertions at the S1/S2 Cleavage Site of the Spike Protein"
.
Current Biology
.
30
(11): 2196?2203.e3.
doi
:
10.1016/j.cub.2020.05.023
.
PMC
7211627
.
PMID
32416074
.
- ^
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, et al. (March 2020).
"A pneumonia outbreak associated with a new coronavirus of probable bat origin"
.
Nature
.
579
(7798): 270?273.
Bibcode
:
2020Natur.579..270Z
.
doi
:
10.1038/s41586-020-2012-7
.
PMC
7095418
.
PMID
32015507
.
- ^
Temmam S, Vongphayloth K, Baquero E, Munier S, Bonomi M, Regnault B, et al. (April 2022). "Bat coronaviruses related to SARS-CoV-2 and infectious for human cells".
Nature
.
604
(7905): 330?336.
Bibcode
:
2022Natur.604..330T
.
doi
:
10.1038/s41586-022-04532-4
.
PMID
35172323
.
S2CID
246902858
.
- ^
Sonnevend, Julia (December 2020). Alexander, Jeffrey C.; Jacobs, Ronald N.; Smith, Philip (eds.).
"A virus as an icon: the 2020 pandemic in images"
(PDF)
.
American Journal of Cultural Sociology
.
8
(3:
The COVID Crisis and Cultural Sociology: Alone Together
).
Basingstoke
:
Palgrave Macmillan
: 451?461.
doi
:
10.1057/s41290-020-00118-7
.
eISSN
2049-7121
.
ISSN
2049-7113
.
PMC
7537773
.
PMID
33042541
.
- ^
Goldsmith CS, Tatti KM, Ksiazek TG, Rollin PE, Comer JA, Lee WW, et al. (February 2004).
"Ultrastructural characterization of SARS coronavirus"
.
Emerging Infectious Diseases
.
10
(2): 320?6.
doi
:
10.3201/eid1002.030913
.
PMC
3322934
.
PMID
15030705
.
Virions acquired an envelope by budding into the cisternae and formed mostly spherical, sometimes pleomorphic, particles that averaged 78 nm in diameter (Figure 1A).
- ^
Neuman BW, Adair BD, Yoshioka C, Quispe JD, Orca G, Kuhn P, et al. (August 2006).
"Supramolecular architecture of severe acute respiratory syndrome coronavirus revealed by electron cryomicroscopy"
.
Journal of Virology
.
80
(16): 7918?28.
doi
:
10.1128/JVI.00645-06
.
PMC
1563832
.
PMID
16873249
.
Particle diameters ranged from 50 to 150 nm, excluding the spikes, with mean particle diameters of 82 to 94 nm; Also See Figure 1 for double shell.
- ^
Lai MM, Cavanagh D (1997).
"The molecular biology of coronaviruses"
.
Advances in Virus Research
.
48
: 1?100.
doi
:
10.1016/S0065-3527(08)60286-9
.
ISBN
9780120398485
.
PMC
7130985
.
PMID
9233431
.
- ^
Masters PS (1 January 2006).
The molecular biology of coronaviruses
. Advances in Virus Research. Vol. 66. Academic Press. pp. 193?292.
doi
:
10.1016/S0065-3527(06)66005-3
.
ISBN
9780120398690
.
PMC
7112330
.
PMID
16877062
.
Nevertheless, the interaction between S protein and receptor remains the principal, if not sole, determinant of coronavirus host species range and tissue tropism.
- ^
Cui J, Li F, Shi ZL (March 2019).
"Origin and evolution of pathogenic coronaviruses"
.
Nature Reviews. Microbiology
.
17
(3): 181?192.
doi
:
10.1038/s41579-018-0118-9
.
PMC
7097006
.
PMID
30531947
.
Different SARS-CoV strains isolated from several hosts vary in their binding affinities for human ACE2 and consequently in their infectivity of human cells76,78 (Fig. 6b)
- ^
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See section: Virion Structure.
- ^
Chang CK, Hou MH, Chang CF, Hsiao CD, Huang TH (March 2014).
"The SARS coronavirus nucleocapsid protein--forms and functions"
.
Antiviral Research
.
103
: 39?50.
doi
:
10.1016/j.antiviral.2013.12.009
.
PMC
7113676
.
PMID
24418573
.
See Figure 4c.
- ^
Neuman BW, Kiss G, Kunding AH, Bhella D, Baksh MF, Connelly S, et al. (April 2011).
"A structural analysis of M protein in coronavirus assembly and morphology"
.
Journal of Structural Biology
.
174
(1): 11?22.
doi
:
10.1016/j.jsb.2010.11.021
.
PMC
4486061
.
PMID
21130884
.
See Figure 10.
- ^
Lal SK, ed. (2010).
Molecular Biology of the SARS-Coronavirus
.
doi
:
10.1007/978-3-642-03683-5
.
ISBN
978-3-642-03682-8
.
- ^
a
b
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See section: Coronavirus Life Cycle ? Attachment and Entry
- ^
a
b
Simmons G, Zmora P, Gierer S, Heurich A, Pohlmann S (December 2013).
"Proteolytic activation of the SARS-coronavirus spike protein: cutting enzymes at the cutting edge of antiviral research"
.
Antiviral Research
.
100
(3): 605?14.
doi
:
10.1016/j.antiviral.2013.09.028
.
PMC
3889862
.
PMID
24121034
.
See Figure 2.
- ^
Starr, Tyler N.; Zepeda, Samantha K.; Walls, Alexandra C.; Greaney, Allison J.; Alkhovsky, Sergey; Veesler, David; Bloom, Jesse D. (1 March 2022).
"ACE2 binding is an ancestral and evolvable trait of sarbecoviruses"
.
Nature
.
603
(7903): 913?918.
Bibcode
:
2022Natur.603..913S
.
doi
:
10.1038/s41586-022-04464-z
.
ISSN
1476-4687
.
PMC
8967715
.
PMID
35114688
.
- ^
Heurich A, Hofmann-Winkler H, Gierer S, Liepold T, Jahn O, Pohlmann S (January 2014).
"TMPRSS2 and ADAM17 cleave ACE2 differentially and only proteolysis by TMPRSS2 augments entry driven by the severe acute respiratory syndrome coronavirus spike protein"
.
Journal of Virology
.
88
(2): 1293?307.
doi
:
10.1128/JVI.02202-13
.
PMC
3911672
.
PMID
24227843
.
The SARS-CoV can hijack two cellular proteolytic systems to ensure the adequate processing of its S protein. Cleavage of SARS-S can be facilitated by cathepsin L, a pH-dependent endo-/lysosomal host cell protease, upon uptake of virions into target cell endosomes (25). Alternatively, the type II transmembrane serine proteases (TTSPs) TMPRSS2 and HAT can activate SARS-S, presumably by cleavage of SARS-S at or close to the cell surface, and activation of SARS-S by TMPRSS2 allows for cathepsin L-independent cellular entry (26,?28).
- ^
Zumla A, Chan JF, Azhar EI, Hui DS, Yuen KY (May 2016).
"Coronaviruses - drug discovery and therapeutic options"
.
Nature Reviews. Drug Discovery
.
15
(5): 327?47.
doi
:
10.1038/nrd.2015.37
.
PMC
7097181
.
PMID
26868298
.
S is activated and cleaved into the S1 and S2 subunits by other host proteases, such as transmembrane protease serine 2 (TMPRSS2) and TMPRSS11D, which enables cell surface non-endosomal virus entry at the plasma membrane.
- ^
Li Z, Tomlinson AC, Wong AH, Zhou D, Desforges M, Talbot PJ, et al. (October 2019).
"The human coronavirus HCoV-229E S-protein structure and receptor binding"
.
eLife
.
8
.
doi
:
10.7554/eLife.51230
.
PMC
6970540
.
PMID
31650956
.
- ^
a
b
c
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See Table 2.
- ^
Rao, S; Hoskins, I; Tonn, T; Garcia, PD; Ozadam, H; Sarinay Cenik, E; Cenik, C (September 2021).
"Genes with 5' terminal oligopyrimidine tracts preferentially escape global suppression of translation by the SARS-CoV-2 Nsp1 protein"
.
RNA
.
27
(9): 1025?1045.
doi
:
10.1261/rna.078661.120
.
PMC
8370740
.
PMID
34127534
.
- ^
Masters PS (1 January 2006).
"The molecular biology of coronaviruses"
.
Advances in Virus Research
.
66
. Academic Press: 193?292.
doi
:
10.1016/S0065-3527(06)66005-3
.
ISBN
9780120398690
.
PMC
7112330
.
PMID
16877062
.
See Figure 8.
- ^
a
b
c
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See section: Replicase Protein Expression
- ^
Mehdi Moustaqil (5 June 2020).
"SARS-CoV-2 proteases cleave IRF3 and critical modulators of inflammatory pathways (NLRP12 and TAB1): implications for disease presentation across species and the search for reservoir hosts"
.
bioRxiv
: 2020.06.05.135699.
doi
:
10.1101/2020.06.05.135699
.
S2CID
219604020
.
- ^
Sexton NR, Smith EC, Blanc H, Vignuzzi M, Peersen OB, Denison MR (August 2016).
"Homology-Based Identification of a Mutation in the Coronavirus RNA-Dependent RNA Polymerase That Confers Resistance to Multiple Mutagens"
.
Journal of Virology
.
90
(16): 7415?28.
doi
:
10.1128/JVI.00080-16
.
PMC
4984655
.
PMID
27279608
.
Finally, these results, combined with those from previous work (33, 44), suggest that CoVs encode at least three proteins involved in fidelity (nsp12-RdRp, nsp14-ExoN, and nsp10), supporting the assembly of a multiprotein replicase-fidelity complex, as described previously (38).
- ^
a
b
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See section: Corona Life Cycle ? Replication and Transcription
- ^
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See Figure 1.
- ^
a
b
Zhang XW, Yap YL, Danchin A. Testing the hypothesis of a recombinant origin of the SARS-associated coronavirus. Arch Virol. 2005 Jan;150(1):1-20. Epub 2004 Oct 11. PMID 15480857
- ^
Stanhope MJ, Brown JR, Amrine-Madsen H. Evidence from the evolutionary analysis of nucleotide sequences for a recombinant history of SARS-CoV. Infect Genet Evol. 2004 Mar;4(1):15-9. PMID 15019585
- ^
a
b
Fehr AR, Perlman S (2015). "Coronaviruses: An Overview of Their Replication and Pathogenesis". In Maier HJ, Bickerton E, Britton P (eds.).
Coronaviruses
. Methods in Molecular Biology. Vol. 1282. Springer. pp. 1?23.
doi
:
10.1007/978-1-4939-2438-7_1
.
ISBN
978-1-4939-2438-7
.
PMC
4369385
.
PMID
25720466
.
See section: Coronavirus Life Cycle ? Assembly and Release
Further reading
[
edit
]
- Peiris JS, Lai ST, Poon LL, Guan Y, Yam LY, Lim W, et al. (April 2003).
"Coronavirus as a possible cause of severe acute respiratory syndrome"
.
Lancet
.
361
(9366): 1319?25.
doi
:
10.1016/s0140-6736(03)13077-2
.
PMC
7112372
.
PMID
12711465
.
- Rota PA, Oberste MS, Monroe SS, Nix WA, Campagnoli R, Icenogle JP, et al. (May 2003).
"Characterization of a novel coronavirus associated with severe acute respiratory syndrome"
.
Science
.
300
(5624): 1394?9.
Bibcode
:
2003Sci...300.1394R
.
doi
:
10.1126/science.1085952
.
PMID
12730500
.
- Marra MA, Jones SJ, Astell CR, Holt RA, Brooks-Wilson A, Butterfield YS, et al. (May 2003).
"The Genome sequence of the SARS-associated coronavirus"
.
Science
.
300
(5624): 1399?404.
Bibcode
:
2003Sci...300.1399M
.
doi
:
10.1126/science.1085953
.
PMID
12730501
.
- Snijder EJ, Bredenbeek PJ, Dobbe JC, Thiel V, Ziebuhr J, Poon LL, et al. (August 2003).
"Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage"
.
Journal of Molecular Biology
.
331
(5): 991?1004.
CiteSeerX
10.1.1.319.7007
.
doi
:
10.1016/S0022-2836(03)00865-9
.
PMC
7159028
.
PMID
12927536
.
S2CID
14974326
.
- Yount B, Roberts RS, Lindesmith L, Baric RS (August 2006).
"Rewiring the severe acute respiratory syndrome coronavirus (SARS-CoV) transcription circuit: engineering a recombination-resistant genome"
.
Proceedings of the National Academy of Sciences of the United States of America
.
103
(33): 12546?51.
Bibcode
:
2006PNAS..10312546Y
.
doi
:
10.1073/pnas.0605438103
.
PMC
1531645
.
PMID
16891412
.
- Thiel V, ed. (2007).
Coronaviruses: Molecular and Cellular Biology
(1st ed.). Caister Academic Press.
ISBN
978-1-904455-16-5
.
- Enjuanes L, Sola I, Zuniga S, Almazan F (2008). "Coronavirus Replication and Interaction with Host". In Mettenleiter TC, Sobrino F (eds.).
Animal Viruses: Molecular Biology
. Caister Academic Press.
ISBN
978-1-904455-22-6
.
External links
[
edit
]
|
|---|
|
| General topics
| |
|---|
| People
| |
|---|
| In popular culture
| |
|---|
| Related articles
| |
|---|
|
|---|
| Severe acute respiratory syndrome-related coronavirus
| |
|---|