Family of RNA viruses including the influenza viruses
This article is about the virus family that contains seven genera. For specific information about the subfamily that affects humans, see
Influenza
.
Orthomyxoviridae
(from
Greek
?ρθ??,
orthos
'straight' + μ?ξα,
myxa
'
mucus
')
[1]
is a family of
negative-sense
RNA viruses
. It includes seven
genera
:
Alphainfluenzavirus
,
Betainfluenzavirus
,
Gammainfluenzavirus
,
Deltainfluenzavirus
,
Isavirus
,
Thogotovirus
, and
Quaranjavirus
. The first four genera contain viruses that cause
influenza
in
birds
(see also
avian influenza
) and
mammals
, including humans. Isaviruses infect
salmon
; the thogotoviruses are
arboviruses
, infecting
vertebrates
and
invertebrates
(such as
ticks
and
mosquitoes
).
[2]
[3]
[4]
The Quaranjaviruses are also
arboviruses
, infecting vertebrates (birds) and invertebrates (
arthropods
).
The four genera of Influenza virus that infect vertebrates, which are identified by antigenic differences in their
nucleoprotein
and
matrix protein
, are as follows:
Structure
[
edit
]
 Influenza A virus structure
Influenza A virus structure
The influenzavirus
virion
is
pleomorphic
; the
viral envelope
can occur in spherical and filamentous forms. In general, the virus's morphology is ellipsoidal with particles 100?120
nm
in diameter, or filamentous with particles 80?100 nm in diameter and up to 20 μm long.
[5]
There are approximately 500 distinct spike-like surface projections in the envelope each projecting 10?14 nm from the surface with varying surface densities. The major
glycoprotein
(HA) spike is interposed irregularly by clusters of
neuraminidase
(NA) spikes, with a ratio of HA to NA of about 10 to 1.
[6]
The viral envelope composed of a
lipid bilayer
membrane in which the glycoprotein spikes are anchored encloses the
nucleocapsids
; nucleoproteins of different size classes with a loop at each end; the arrangement within the virion is uncertain. The ribonuclear proteins are filamentous and fall in the range of 50?130 nm long and 9?15 nm in diameter with helical symmetry.
[
citation needed
]
Genome
[
edit
]
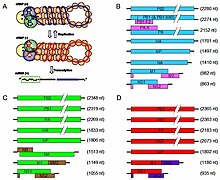 Influenzavirus genomes. Segments translate to
polymerase
(PB1, PB2, and PA),
hemagglutinin
(HA),
neuramindase
(NA),
nucleoprotein
(NP), membrane protein (M), and
non-structural protein
(NS).
Influenzavirus genomes. Segments translate to
polymerase
(PB1, PB2, and PA),
hemagglutinin
(HA),
neuramindase
(NA),
nucleoprotein
(NP), membrane protein (M), and
non-structural protein
(NS).
Viruses of the family
Orthomyxoviridae
contain six to eight segments of linear
negative-sense
single stranded RNA. They have a total genome length that is 10,000?14,600
nucleotides
(nt).
[7]
The influenza A
genome
, for instance, has eight pieces of segmented negative-sense
RNA
(13.5 kilobases total).
[8]
The best-characterised of the influenzavirus proteins are
hemagglutinin
and
neuraminidase
, two large
glycoproteins
found on the outside of the viral particles. Hemagglutinin is a
lectin
that mediates binding of the virus to target cells and entry of the viral genome into the target cell.
[9]
In contrast, neuraminidase is an
enzyme
involved in the release of
progeny
virus from infected cells, by cleaving sugars that bind the mature viral particles. The hemagglutinin (H) and neuraminidase (N)
proteins
are key targets for antibodies and antiviral drugs,
[10]
[11]
and they are used to classify the different
serotypes
of influenza A viruses, hence the
H
and
N
in
H5N1
.
The genome sequence has terminal repeated sequences; repeated at both ends. Terminal repeats at the 5′-end 12?13 nucleotides long. Nucleotide sequences of 3′-terminus identical; the same in genera of same family; most on RNA (segments), or on all RNA species. Terminal repeats at the 3′-end 9?11 nucleotides long. Encapsidated nucleic acid is solely genomic. Each virion may contain defective interfering copies. In Influenza A (H1N1) PB1-F2 is produced from an alternative reading frame in PB1. The M and NS genes produce two different genes via
alternative splicing
.
[12]
Replication cycle
[
edit
]
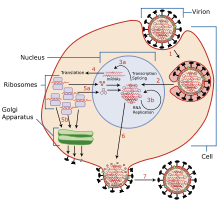 Infection and replication of the influenza virus. The steps in this process are discussed in the text.
Infection and replication of the influenza virus. The steps in this process are discussed in the text.
Typically, influenza is transmitted from infected mammals through the air by coughs or sneezes, creating
aerosols
containing the virus, and from infected birds through their
droppings
. Influenza can also be transmitted by
saliva
,
nasal secretions
,
feces
and
blood
. Infections occur through contact with these bodily fluids or with contaminated surfaces. Out of a host, flu viruses can remain infectious for about one week at human body temperature, over 30 days at 0 °C (32 °F), and indefinitely at very low temperatures (such as lakes in northeast
Siberia
). They can be inactivated easily by
disinfectants
and
detergents
.
[13]
[14]
[15]
The viruses bind to a cell through interactions between its
hemagglutinin
glycoprotein and
sialic acid
sugars on the surfaces of
epithelial cells
in the lung and throat (Stage 1 in infection figure).
[16]
The cell imports the virus by
endocytosis
. In the acidic
endosome
, part of the hemagglutinin protein fuses the viral envelope with the vacuole's membrane, releasing the viral RNA (vRNA) molecules, accessory proteins and
RNA-dependent RNA polymerase
into the
cytoplasm
(Stage 2).
[17]
These proteins and vRNA form a complex that is transported into the
cell nucleus
, where the RNA-dependent RNA polymerase begins transcribing complementary positive-sense cRNA (Steps 3a and b).
[18]
The cRNA is either exported into the cytoplasm and translated (step 4), or remains in the nucleus. Newly synthesised viral proteins are either secreted through the
Golgi apparatus
onto the cell surface (in the case of neuraminidase and hemagglutinin, step 5b) or transported back into the nucleus to bind vRNA and form new viral genome particles (step 5a). Other viral proteins have multiple actions in the host cell, including degrading cellular
mRNA
and using the released
nucleotides
for vRNA synthesis and also inhibiting translation of host-cell mRNAs.
[19]
Negative-sense vRNAs that form the
genomes
of future viruses, RNA-dependent RNA transcriptase, and other viral proteins are assembled into a virion. Hemagglutinin and neuraminidase molecules cluster into a bulge in the cell membrane. The vRNA and viral core proteins leave the nucleus and enter this membrane protrusion (step 6). The mature virus buds off from the cell in a sphere of host phospholipid membrane, acquiring hemagglutinin and neuraminidase with this membrane coat (step 7).
[20]
As before, the viruses adhere to the cell through hemagglutinin; the mature viruses detach once their
neuraminidase
has cleaved sialic acid residues from the host cell.
[16]
After the release of new influenza virus, the host cell dies.
 Transcription of mRNAs initiated by viral polymerase using
cap snatching
Transcription of mRNAs initiated by viral polymerase using
cap snatching
Orthomyxoviridae viruses are one of two RNA viruses that replicate in the nucleus (the other being
retroviridae
). This is because the machinery of orthomyxo viruses cannot make their own mRNAs. They use cellular RNAs as primers for initiating the viral mRNA synthesis in a process known as
cap snatching
.
[21]
Once in the nucleus, the RNA Polymerase Protein PB2 finds a cellular pre-mRNA and binds to its 5′ capped end. Then RNA Polymerase PA cleaves off the cellular mRNA near the 5′ end and uses this capped fragment as a primer for transcribing the rest of the viral RNA genome in viral mRNA.
[22]
This is due to the need of mRNA to have a 5′ cap in order to be recognized by the cell's
ribosome
for translation.
Since RNA
proofreading
enzymes are absent, the RNA-dependent RNA transcriptase makes a single nucleotide insertion error roughly every 10 thousand nucleotides, which is the approximate length of the influenza vRNA. Hence, nearly every newly manufactured influenza virus will contain a mutation in its genome.
[23]
The separation of the genome into eight separate segments of vRNA allows mixing (
reassortment
) of the genes if more than one variety of influenza virus has infected the same cell (
superinfection
). The resulting alteration in the genome segments packaged into viral progeny confers new behavior, sometimes the ability to infect new host species or to overcome protective immunity of host populations to its old genome (in which case it is called an
antigenic shift
).
[10]
Classification
[
edit
]
In a
phylogenetic
-based
taxonomy
, the category
RNA virus
includes the subcategory
negative-sense ssRNA virus
, which includes the order
Articulavirales
, and the family
Orthomyxoviridae
. The genera-associated species and
serotypes
of
Orthomyxoviridae
are shown in the following table.
Orthomyxovirus Genera, Species, and Serotypes
| Genus
|
Species
(* indicates
type species
)
|
Serotypes
or Subtypes
|
Hosts
|
| Alphainfluenzavirus
|
Influenza A virus
*
|
H1N1
,
H1N2
,
H2N2
,
H3N1
,
H3N2
,
H3N8
,
H5N1
,
H5N2
,
H5N3
,
H5N8
,
H5N9
,
H7N1
,
H7N2
,
H7N3
,
H7N4
,
H7N7
,
H7N9
,
H9N2
,
H10N7
|
Human
,
pig
,
bird
,
horse
,
bat
|
| Betainfluenzavirus
|
Influenza B virus
*
|
Victoria, Yamagata
[24]
|
Human,
seal
|
| Gammainfluenzavirus
|
Influenza C virus
*
|
|
Human, pig
|
| Deltainfluenzavirus
|
Influenza D virus
*
|
|
Pig, cattle
|
| Isavirus
|
Infectious salmon anemia virus
*
|
|
Atlantic salmon
|
| Thogotovirus
|
Thogotovirus
*
|
|
Tick
,
mosquito
,
mammal
(including human)
|
| Dhori virus
|
Batken virus
,
Bourbon virus
,
Jos virus
|
| Quaranjavirus
[25]
|
| Quaranfil virus
,*
Johnston Atoll virus
|
|
|
Types
[
edit
]
There are four genera of influenza virus, each containing only a single species, or type. Influenza A and C infect a variety of species (including humans), while influenza B almost exclusively infects humans, and influenza D infects cattle and pigs.
[26]
[27]
[28]
Influenza A
[
edit
]
 Diagram of influenza nomenclature
Diagram of influenza nomenclature
Influenza A viruses are further classified, based on the viral surface proteins
hemagglutinin
(HA or H) and
neuraminidase
(NA or N). 18 HA subtypes (or serotypes) and 11 NA subtypes of influenza A virus have been isolated in nature. Among these, the HA subtype 1-16 and NA subtype 1-9 are found in wild waterfowl and shorebirds and the HA subtypes 17-18 and NA subtypes 10-11 have only been isolated from bats.
[29]
[30]
Further variation exists; thus, specific influenza strain
isolates
are identified by the
Influenza virus nomenclature,
[31]
specifying virus type, host species (if not human), geographical location where first isolated, laboratory reference, year of isolation, and HA and NA subtype.
[32]
[33]
Examples of the nomenclature are:
- A/Brisbane/59/2007 (H1N1)
- isolated from a human
- A/swine/South Dakota/152B/2009 (H1N2)
- isolated from a pig
The type A influenza viruses are the most virulent human pathogens among the three influenza types and cause the most severe disease. It is thought that all influenza A viruses causing outbreaks or pandemics originate from wild aquatic birds.
[34]
All influenza A virus pandemics since the 1900s were caused by
Avian influenza
, through
Reassortment
with other influenza strains, either those that affect humans (seasonal flu) or those affecting other animals (see
2009 swine flu pandemic
).
[35]
The serotypes that have been confirmed in
humans
, ordered by the number of confirmed human deaths, are:
Influenza B
[
edit
]
 Host range of influenza viruses
Host range of influenza viruses
Influenza B virus is almost exclusively a human pathogen, and is less common than influenza A. The only other animal known to be susceptible to influenza B infection is the
seal
.
[47]
This type of influenza mutates at a rate 2?3 times lower than type A
[48]
and consequently is less genetically diverse, with only one influenza B serotype.
[26]
As a result of this lack of
antigenic
diversity, a degree of immunity to influenza B is usually acquired at an early age. However, influenza B mutates enough that lasting immunity is not possible.
[49]
This reduced rate of antigenic change, combined with its limited host range (inhibiting cross species
antigenic shift
), ensures that pandemics of influenza B do not occur.
[50]
Influenza C
[
edit
]
The influenza C virus infects
humans
and
pigs
, and can cause severe illness and local
epidemics
.
[51]
However, influenza C is less common than the other types and usually causes mild disease in children.
[52]
[53]
Influenza D
[
edit
]
This is a genus that was classified in 2016, the members of which were first isolated in 2011.
[54]
This genus appears to be most closely related to Influenza C, from which it diverged several hundred years ago.
[55]
There are at least two extant strains of this genus.
[56]
The main hosts appear to be cattle, but the virus has been known to infect pigs as well.
Viability and disinfection
[
edit
]
Mammalian influenza viruses tend to be labile, but can survive several hours in mucus.
[57]
Avian influenza virus can survive for 100 days in distilled water at room temperature, and 200 days at 17 °C (63 °F). The avian virus is inactivated more quickly in manure, but can survive for up to two weeks in feces on cages. Avian influenza viruses can survive indefinitely when frozen.
[57]
Influenza viruses are susceptible to bleach, 70% ethanol, aldehydes, oxidizing agents, and quaternary ammonium compounds. They are inactivated by heat of 133 °F (56 °C) for minimum of 60 minutes, as well as by low pH <2.
[57]
Vaccination and prophylaxis
[
edit
]
 Targets of anti-influenza agents that are licensed or under investigation
Targets of anti-influenza agents that are licensed or under investigation
Vaccines and drugs are available for the prophylaxis and treatment of influenza virus infections. Vaccines are composed of either inactivated or live attenuated virions of the H1N1 and H3N2 human influenza A viruses, as well as those of influenza B viruses. Because the antigenicities of the wild viruses evolve, vaccines are reformulated annually by updating the seed strains.
[
citation needed
]
When the antigenicities of the seed strains and wild viruses do not match, vaccines fail to protect the vaccinees.
[
citation needed
]
In addition, even when they do match, escape mutants are often generated.
[
citation needed
]
Drugs available for the treatment of influenza include
Amantadine
and
Rimantadine
, which inhibit the uncoating of virions by interfering with M2, and
Oseltamivir
(marketed under the brand name
Tamiflu
),
Zanamivir
, and
Peramivir
, which inhibit the release of virions from infected cells by interfering with NA. However, escape mutants are often generated for the former drug and less frequently for the latter drug.
[58]
See also
[
edit
]
References
[
edit
]
- ^
International Committee on Taxonomy of Viruses
Index of Viruses ? Orthomyxovirus (2006). In: ICTVdB?The Universal Virus Database, version 4. Buchen-Osmond, C (Ed), Columbia University, New York.
- ^
Jones LD, Nuttall PA (1989). "Non-viraemic transmission of Thogoto virus: influence of time and distance".
Trans. R. Soc. Trop. Med. Hyg
.
83
(5): 712?14.
doi
:
10.1016/0035-9203(89)90405-7
.
PMID
2617637
.
- ^
Ely B (1999).
"Infectious Salmon Anaemia"
.
Mill Hill Essays
.
National Institute for Medical Research
. Archived from
the original
on 2007-08-24
. Retrieved
2007-09-14
.
- ^
Raynard RS, Murray AG, Gregory A (2001).
"Infectious salmon anaemia virus in wild fish from Scotland"
.
Dis. Aquat. Org
.
46
(2): 93?100.
doi
:
10.3354/dao046093
.
PMID
11678233
.
- ^
Noda T (2012-01-03).
"Native morphology of influenza virions"
.
Frontiers in Microbiology
.
2
: 269.
doi
:
10.3389/fmicb.2011.00269
.
PMC
3249889
.
PMID
22291683
.
- ^
Einav T, Gentles LE, Bloom JD (2020-07-23).
"SnapShot: Influenza by the Numbers"
(PDF)
.
Cell
.
182
(2): 532?532.e1.
doi
:
10.1016/j.cell.2020.05.004
.
PMID
32707094
.
S2CID
220715148
.
- ^
"ICTV Ninth Report; 2009 Taxonomy Release:
Orthomyxoviridae
"
.
ICTV
. Retrieved
19 September
2020
.
- ^
Ghedin E, Sengamalay NA, Shumway M, Zaborsky J, Feldblyum T, Subbu V, Spiro DJ, Sitz J, Koo H, Bolotov P, Dernovoy D, Tatusova T, Bao Y, St George K, Taylor J, Lipman DJ, Fraser CM, Taubenberger JK, Salzberg SL (October 2005).
"Large-scale sequencing of human influenza reveals the dynamic nature of viral genome evolution"
.
Nature
.
437
(7062): 1162?6.
Bibcode
:
2005Natur.437.1162G
.
doi
:
10.1038/nature04239
.
PMID
16208317
.
- ^
Suzuki Y (March 2005).
"Sialobiology of influenza: molecular mechanism of host range variation of influenza viruses"
.
Biological & Pharmaceutical Bulletin
.
28
(3): 399?408.
doi
:
10.1248/bpb.28.399
.
PMID
15744059
.
- ^
a
b
c
Hilleman MR (August 2002). "Realities and enigmas of human viral influenza: pathogenesis, epidemiology and control".
Vaccine
.
20
(25?26): 3068?87.
doi
:
10.1016/S0264-410X(02)00254-2
.
PMID
12163258
.
- ^
Wilson JC, von Itzstein M (July 2003). "Recent strategies in the search for new anti-influenza therapies".
Current Drug Targets
.
4
(5): 389?408.
doi
:
10.2174/1389450033491019
.
PMID
12816348
.
- ^
Bouvier NM, Palese P (September 2008).
"The biology of influenza viruses"
.
Vaccine
.
26
(Suppl 4): D49?53.
doi
:
10.1016/j.vaccine.2008.07.039
.
PMC
3074182
.
PMID
19230160
.
- ^
Suarez DL, Spackman E, Senne DA, Bulaga L, Welsch AC, Froberg K (2003). "The effect of various disinfectants on detection of avian influenza virus by real time RT-PCR".
Avian Diseases
.
47
(3 Suppl): 1091?5.
doi
:
10.1637/0005-2086-47.s3.1091
.
PMID
14575118
.
S2CID
8612187
.
- ^
"Avian Influenza (Bird Flu) Implications for Human Disease. Physical characteristics of influenza A viruses"
.
CIDRAP - Center for Infectious Disease Research and Policy
. University of Minnesota. 12 March 2024.
- ^
"Flu viruses 'can live for decades' on ice"
.
The New Zealand Herald
.
Reuters
. November 30, 2006
. Retrieved
November 1,
2011
.
- ^
a
b
Wagner R, Matrosovich M, Klenk H (May?Jun 2002). "Functional balance between haemagglutinin and neuraminidase in influenza virus infections".
Rev Med Virol
.
12
(3): 159?66.
doi
:
10.1002/rmv.352
.
PMID
11987141
.
S2CID
30876482
.
- ^
Lakadamyali M, Rust M, Babcock H, Zhuang X (Aug 5, 2003).
"Visualizing infection of individual influenza viruses"
.
Proc Natl Acad Sci USA
.
100
(16): 9280?85.
Bibcode
:
2003PNAS..100.9280L
.
doi
:
10.1073/pnas.0832269100
.
PMC
170909
.
PMID
12883000
.
- ^
Cros J, Palese P (September 2003). "Trafficking of viral genomic RNA into and out of the nucleus: influenza, Thogoto and Borna disease viruses".
Virus Res
.
95
(1?2): 3?12.
doi
:
10.1016/S0168-1702(03)00159-X
.
PMID
12921991
.
- ^
Kash J, Goodman A, Korth M, Katze M (July 2006). "Hijacking of the host-cell response and translational control during influenza virus infection".
Virus Res
.
119
(1): 111?20.
doi
:
10.1016/j.virusres.2005.10.013
.
PMID
16630668
.
- ^
Nayak D, Hui E, Barman S (December 2004).
"Assembly and budding of influenza virus"
.
Virus Res
.
106
(2): 147?65.
doi
:
10.1016/j.virusres.2004.08.012
.
PMC
7172797
.
PMID
15567494
.
- ^
"Cap Snatching"
.
ViralZone
. expasy
. Retrieved
11 September
2014
.
- ^
Dias A, Bouvier D, Crepin T, McCarthy AA, Hart DJ, Baudin F, Cusack S, Ruigrok RW (April 2009). "The cap-snatching endonuclease of influenza virus polymerase resides in the PA subunit".
Nature
.
458
(7240): 914?8.
Bibcode
:
2009Natur.458..914D
.
doi
:
10.1038/nature07745
.
PMID
19194459
.
S2CID
4421958
.
- ^
Drake J (May 1, 1993).
"Rates of spontaneous mutation among RNA viruses"
.
Proc Natl Acad Sci USA
.
90
(9): 4171?5.
Bibcode
:
1993PNAS...90.4171D
.
doi
:
10.1073/pnas.90.9.4171
.
PMC
46468
.
PMID
8387212
.
- ^
Biere B, Bauer B, Schweiger B (April 2010).
"Differentiation of influenza B virus lineages Yamagata and Victoria by real-time PCR"
(PDF)
.
Journal of Clinical Microbiology
.
48
(4): 1425?7.
doi
:
10.1128/JCM.02116-09
.
PMC
2849545
.
PMID
20107085
.
- ^
ICTV Taxonomy History
, ICTV, 2014, archived from
the original
on 2 April 2015
, retrieved
6 June
2006
- ^
a
b
Hay A, Gregory V, Douglas A, Lin Y (Dec 29, 2001).
"The evolution of human influenza viruses"
.
Philos Trans R Soc Lond B Biol Sci
.
356
(1416): 1861?70.
doi
:
10.1098/rstb.2001.0999
.
PMC
1088562
.
PMID
11779385
.
- ^
"Avian Influenza (Bird Flu)"
. Centers for Disease Control and Prevention
. Retrieved
2007-09-15
.
- ^
Kumar, Binod; Asha, Kumari; Khanna, Madhu; Ronsard, Larance; Meseko, Clement Adebajo; Sanicas, Melvin (April 2018).
"The emerging influenza virus threat: status and new prospects for its therapy and control"
.
Archives of Virology
.
163
(4): 831?844.
doi
:
10.1007/s00705-018-3708-y
.
ISSN
1432-8798
.
PMC
7087104
.
PMID
29322273
.
- ^
Alexander, Dennis J. (2007-07-26).
"An overview of the epidemiology of avian influenza"
.
Vaccine
.
25
(30): 5637?5644.
doi
:
10.1016/j.vaccine.2006.10.051
.
ISSN
0264-410X
.
PMID
17126960
.
- ^
Wu, Ying; Wu, Yan; Tefsen, Boris; Shi, Yi; Gao, George F. (April 2014).
"Bat-derived influenza-like viruses H17N10 and H18N11"
.
Trends in Microbiology
.
22
(4): 183?191.
doi
:
10.1016/j.tim.2014.01.010
.
ISSN
0966-842X
.
PMC
7127364
.
PMID
24582528
.
- ^
"A revision of the system of nomenclature for influenza viruses: a WHO Memorandum"
.
Bull World Health Organ
.
58
(4): 585?591. 1980.
PMC
2395936
.
PMID
6969132
.
This Memorandum was drafted by the signatories listed on page 590 on the occasion of a meeting held in Geneva in February 1980.
- ^
Atkinson W, Hamborsky J, McIntyre L, Wolfe S, eds. (2007).
Epidemiology and Prevention of Vaccine-Preventable Diseases
(10th ed.). Washington DC: Centers for Disease Control and Prevention.
- ^
"Avian Influenza (Bird Flu): Implications for Human Disease"
. Center for Infectious Disease Research & Policy,
University of Minnesota
. 2007-06-27
. Retrieved
2007-09-14
.
- ^
Webster, R. G.; Bean, W. J.; Gorman, O. T.; Chambers, T. M.; Kawaoka, Y. (March 1992).
"Evolution and ecology of influenza A viruses"
.
Microbiological Reviews
.
56
(1): 152?179.
doi
:
10.1128/mr.56.1.152-179.1992
.
ISSN
0146-0749
.
PMC
372859
.
PMID
1579108
.
- ^
Taubenberger, Jeffery K.; Morens, David M. (April 2010).
"Influenza: the once and future pandemic"
.
Public Health Reports
.
125
(Suppl 3): 16?26.
doi
:
10.1177/00333549101250S305
.
ISSN
0033-3549
.
PMC
2862331
.
PMID
20568566
.
- ^
Wang TT,
Palese P
(June 2009).
"Unraveling the Mystery of Swine Influenza Virus"
.
Cell
.
137
(6): 983?85.
doi
:
10.1016/j.cell.2009.05.032
.
PMID
19524497
.
- ^
Taubenberger, JK, Morens, DM (April 2009).
"Pandemic influenza ? including a risk assessment of H5N1"
.
Rev. Sci. Tech. Off. Int. Epiz
.
28
(1): 187?202.
doi
:
10.20506/rst.28.1.1879
.
PMC
2720801
.
PMID
19618626
.
- ^
Fouchier R, Schneeberger P, Rozendaal F, Broekman J, Kemink S, Munster V, Kuiken T, Rimmelzwaan G, Schutten M, Van Doornum G, Koch G, Bosman A, Koopmans M, Osterhaus A (2004).
"Avian influenza A virus (H7N7) associated with human conjunctivitis and a fatal case of acute respiratory distress syndrome"
.
Proc Natl Acad Sci USA
.
101
(5): 1356?61.
Bibcode
:
2004PNAS..101.1356F
.
doi
:
10.1073/pnas.0308352100
.
PMC
337057
.
PMID
14745020
.
- ^
Malik-Peiris JS, Poon LL, Guan Y (July 2009).
"Emergence of a novel swine-origin influenza A virus (S-OIV) H1N1 virus in humans"
.
J Clin Virol
.
45
(3): 169?173.
doi
:
10.1016/j.jcv.2009.06.006
.
PMC
4894826
.
PMID
19540800
.
- ^
Potter CW (October 2001). "A history of influenza".
Journal of Applied Microbiology
.
91
(4): 572?9.
doi
:
10.1046/j.1365-2672.2001.01492.x
.
PMID
11576290
.
S2CID
26392163
.
- ^
"Ten things you need to know about pandemic influenza"
. World Health Organization. 14 October 2005. Archived from
the original
on 23 September 2009
. Retrieved
26 September
2009
.
- ^
Valleron AJ, Cori A, Valtat S, Meurisse S, Carrat F, Boelle PY (May 2010).
"Transmissibility and geographic spread of the 1889 influenza pandemic"
.
Proc. Natl. Acad. Sci. USA
.
107
(19): 8778?81.
Bibcode
:
2010PNAS..107.8778V
.
doi
:
10.1073/pnas.1000886107
.
PMC
2889325
.
PMID
20421481
.
- ^
Mills CE,
Robins JM
, Lipsitch M (December 2004).
"Transmissibility of 1918 pandemic influenza"
.
Nature
.
432
(7019): 904?06.
Bibcode
:
2004Natur.432..904M
.
doi
:
10.1038/nature03063
.
PMC
7095078
.
PMID
15602562
.
- ^
Donaldson LJ, Rutter PD, Ellis BM, et al. (2009).
"Mortality from pandemic A/H1N1 2009 influenza in England: public health surveillance study"
.
BMJ
.
339
: b5213.
doi
:
10.1136/bmj.b5213
.
PMC
2791802
.
PMID
20007665
.
- ^
"ECDC Daily Update ? Pandemic (H1N1) 2009 ? January 18, 2010"
(PDF)
.
European Centre for Disease Prevention and Control
. 2010-01-18. Archived from
the original
(PDF)
on January 22, 2010
. Retrieved
2010-01-18
.
- ^
Dawood FS, Iuliano AD, Reed C, Meltzer MI, Shay DK, Cheng PY, Bandaranayake D, Breiman RF, Brooks WA, Buchy P, Feikin DR, Fowler KB, Gordon A, Hien NT, Horby P, Huang QS, Katz MA, Krishnan A, Lal R, Montgomery JM, Mølbak K, Pebody R, Presanis AM, Razuri H, Steens A, Tinoco YO, Wallinga J, Yu H, Vong S, Bresee J, Widdowson MA (September 2012).
"Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: a modelling study"
.
The Lancet. Infectious Diseases
(Submitted manuscript).
12
(9): 687?95.
doi
:
10.1016/S1473-3099(12)70121-4
.
PMID
22738893
.
Archived
(PDF)
from the original on Apr 9, 2024 – via Zenodo.
- ^
Osterhaus AD, Rimmelzwaan GF, Martina BE, Bestebroer TM, Fouchier RA (May 2000). "Influenza B virus in seals".
Science
.
288
(5468): 1051?3.
Bibcode
:
2000Sci...288.1051O
.
doi
:
10.1126/science.288.5468.1051
.
PMID
10807575
.
- ^
Nobusawa E, Sato K (April 2006).
"Comparison of the mutation rates of human influenza A and B viruses"
.
Journal of Virology
.
80
(7): 3675?8.
doi
:
10.1128/JVI.80.7.3675-3678.2006
.
PMC
1440390
.
PMID
16537638
.
- ^
Webster RG, Bean WJ, Gorman OT, Chambers TM, Kawaoka Y (March 1992).
"Evolution and ecology of influenza A viruses"
.
Microbiological Reviews
.
56
(1): 152?79.
doi
:
10.1128/MMBR.56.1.152-179.1992
.
PMC
372859
.
PMID
1579108
.
- ^
Zambon MC (November 1999).
"Epidemiology and pathogenesis of influenza"
.
The Journal of Antimicrobial Chemotherapy
. 44 Suppl B (Suppl B): 3?9.
doi
:
10.1093/jac/44.suppl_2.3
.
PMID
10877456
.
S2CID
15968981
.
- ^
Matsuzaki Y, Sugawara K, Mizuta K, Tsuchiya E, Muraki Y, Hongo S, Suzuki H, Nakamura K (2002).
"Antigenic and genetic characterization of influenza C viruses which caused two outbreaks in Yamagata City, Japan, in 1996 and 1998"
.
J Clin Microbiol
.
40
(2): 422?29.
doi
:
10.1128/JCM.40.2.422-429.2002
.
PMC
153379
.
PMID
11825952
.
- ^
Matsuzaki Y, Katsushima N, Nagai Y, Shoji M, Itagaki T, Sakamoto M, Kitaoka S, Mizuta K, Nishimura H (May 1, 2006).
"Clinical features of influenza C virus infection in children"
.
J Infect Dis
.
193
(9): 1229?35.
doi
:
10.1086/502973
.
PMID
16586359
.
- ^
Katagiri S, Ohizumi A, Homma M (July 1983). "An outbreak of type C influenza in a children's home".
J Infect Dis
.
148
(1): 51?56.
doi
:
10.1093/infdis/148.1.51
.
PMID
6309999
.
- ^
Hause BM, Ducatez M, Collin EA, Ran Z, Liu R, Sheng Z, Armien A, Kaplan B, Chakravarty S, Hoppe AD, Webby RJ, Simonson RR, Li F (February 2013).
"Isolation of a novel swine influenza virus from Oklahoma in 2011 which is distantly related to human influenza C viruses"
.
PLOS Pathogens
.
9
(2): e1003176.
doi
:
10.1371/journal.ppat.1003176
.
PMC
3567177
.
PMID
23408893
.
- ^
Sheng Z, Ran Z, Wang D, Hoppe AD, Simonson R, Chakravarty S, Hause BM, Li F (February 2014).
"Genomic and evolutionary characterization of a novel influenza-C-like virus from swine"
.
Archives of Virology
.
159
(2): 249?55.
doi
:
10.1007/s00705-013-1815-3
.
PMC
5714291
.
PMID
23942954
.
- ^
Collin EA, Sheng Z, Lang Y, Ma W, Hause BM, Li F (January 2015).
"Cocirculation of two distinct genetic and antigenic lineages of proposed influenza D virus in cattle"
.
Journal of Virology
.
89
(2): 1036?42.
doi
:
10.1128/JVI.02718-14
.
PMC
4300623
.
PMID
25355894
.
- ^
a
b
c
Spickler AR (February 2016).
"Influenza"
(PDF)
.
The Center for Food Security and Public Health
. Iowa State University. p. 7.
- ^
Suzuki Y (October 2006).
"Natural selection on the influenza virus genome"
.
Molecular Biology and Evolution
.
23
(10). Oxford Academic: 1902?11.
doi
:
10.1093/molbev/msl050
.
PMID
16818477
.
Archived
from the original on Jan 21, 2022.
Further reading
[
edit
]
External links
[
edit
]