Interdisciplinary research area
 A
ribosome
is a
biological machine
that utilizes
protein dynamics
A
ribosome
is a
biological machine
that utilizes
protein dynamics
Molecular biophysics
is a rapidly evolving interdisciplinary area of research that combines concepts in
physics
,
chemistry
,
engineering
,
mathematics
and
biology
.
[1]
It seeks to understand
biomolecular
systems and explain biological function in terms of molecular structure, structural organization, and dynamic behaviour at various levels of complexity (from single molecules to
supramolecular
structures,
viruses
and small living systems). This discipline covers topics such as the measurement of molecular forces, molecular associations,
allosteric interactions
,
Brownian motion
, and
cable theory
.
[2]
Additional areas of study can be found on
Outline of Biophysics
. The discipline has required development of specialized equipment and procedures capable of imaging and manipulating minute living structures, as well as novel experimental approaches.
Overview
[
edit
]
Molecular biophysics typically addresses biological questions similar to those in
biochemistry
and
molecular biology
, seeking to find the physical underpinnings of biomolecular phenomena. Scientists in this field conduct research concerned with understanding the interactions between the various systems of a cell, including the interactions between
DNA
,
RNA
and
protein biosynthesis
, as well as how these interactions are regulated. A great variety of techniques are used to answer these questions.
Fluorescent
imaging techniques, as well as
electron microscopy
,
X-ray crystallography
,
NMR spectroscopy
,
atomic force microscopy
(AFM) and
small-angle scattering
(SAS) both with
X-rays
and
neutrons
(SAXS/SANS) are often used to visualize structures of biological significance.
Protein dynamics
can be observed by
neutron spin echo
spectroscopy.
Conformational change
in structure can be measured using techniques such as
dual polarisation interferometry
,
circular dichroism
,
SAXS
and
SANS
. Direct manipulation of molecules using
optical tweezers
or
AFM
, can also be used to monitor biological events where forces and distances are at the
nanoscale
. Molecular biophysicists often consider complex biological events as systems of interacting entities which can be understood e.g. through
statistical mechanics
,
thermodynamics
and
chemical kinetics
. By drawing knowledge and experimental techniques from a wide variety of disciplines, biophysicists are often able to directly observe, model or even manipulate the structures and interactions of individual
molecules
or complexes of molecules.
Areas of Research
[
edit
]
![[icon]](//upload.wikimedia.org/wikipedia/commons/thumb/1/1c/Wiki_letter_w_cropped.svg/20px-Wiki_letter_w_cropped.svg.png) | This section
needs expansion
. You can help by
adding to it
.
(
June 2019
)
|
Computational biology
[
edit
]
Computational biology involves the development and application of data-analytical and theoretical methods, mathematical modeling and computational simulation techniques to the study of biological, ecological, behavioral, and social systems. The field is broadly defined and includes foundations in biology,
applied mathematics
,
statistics
,
biochemistry
, chemistry,
biophysics
,
molecular biology
,
genetics
,
genomics
,
computer science
and
evolution
. Computational biology has become an important part of developing emerging technologies for the field of biology.
[3]
Molecular modelling encompasses all methods, theoretical and computational, used to
model
or mimic the behaviour of
molecules
. The methods are used in the fields of
computational chemistry
,
drug design
,
computational biology
and
materials science
to study molecular systems ranging from small chemical systems to large biological molecules and material assemblies.
[4]
[5]
Membrane biophysics
[
edit
]
Membrane biophysics is the study of
biological membrane
structure and function using physical,
computational
, mathematical, and
biophysical methods
. A combination of these methods can be used to create
phase diagrams
of different types of membranes, which yields information on
thermodynamic
behavior of a membrane and its components. As opposed to membrane biology, membrane biophysics focuses on quantitative information and modeling of various membrane phenomena, such as
lipid raft
formation, rates of lipid and cholesterol flip-flop, protein-lipid coupling, and the effect of bending and elasticity functions of membranes on inter-cell connections.
[6]
Motor proteins
[
edit
]
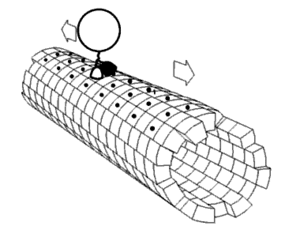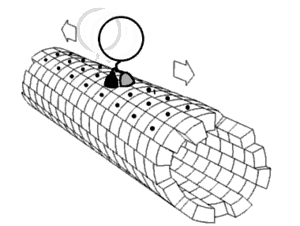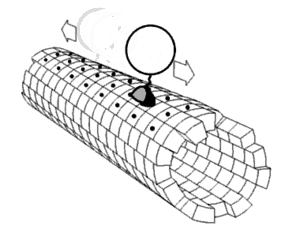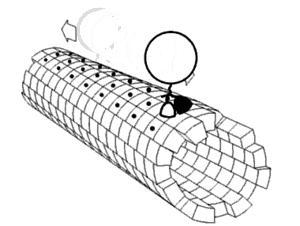 Kinesin
walking on a
microtubule
is a molecular
biological machine
using
protein domain dynamics
on
nanoscales
Kinesin
walking on a
microtubule
is a molecular
biological machine
using
protein domain dynamics
on
nanoscales
Motor proteins are a class of
molecular motors
that can move along the cytoplasm of animal cells. They convert chemical energy into mechanical work by the
hydrolysis
of
ATP
. A good example is the
muscle
protein
myosin
which "motors" the contraction of muscle fibers in animals. Motor proteins are the driving force behind most
active transport
of
proteins
and
vesicles
in the
cytoplasm
.
Kinesins
and
cytoplasmic dyneins
play essential roles in intracellular transport such as
axonal transport
and in the formation of the
spindle apparatus
and the separation of the
chromosomes
during
mitosis
and
meiosis
. Axonemal dynein, found in
cilia
and
flagella
, is crucial to
cell motility
, for example in
spermatozoa
, and fluid transport, for example in trachea.
Some biological machines are motor proteins, such as myosin, which is responsible for
muscle
contraction,
kinesin
, which moves cargo inside cells away from the
nucleus
along
microtubules
, and
dynein
, which moves cargo inside cells towards the nucleus and produces the axonemal beating of
motile cilia
and flagella. "[I]n effect, the [motile cilium] is a nanomachine composed of perhaps over 600 proteins in molecular complexes, many of which also function independently as nanomachines...
Flexible linkers
allow the
mobile protein domains
connected by them to recruit their binding partners and induce long-range
allostery
via
protein domain dynamics
.
[7]
Other biological machines are responsible for energy production, for example
ATP synthase
which harnesses energy from
proton gradients across membranes
to drive a turbine-like motion used to synthesise ATP, the energy currency of a cell.
[8]
Still other machines are responsible for
gene expression
, including
DNA polymerases
for replicating DNA,
RNA polymerases
for producing
mRNA
, the
spliceosome
for removing
introns
, and the
ribosome
for
synthesising proteins
. These machines and their
nanoscale dynamics
are far more complex than any
molecular machines
that have yet been artificially constructed.
[9]
These
molecular motors
are the essential agents of movement in living organisms. In general terms, a
motor
is a device that consumes energy in one form and converts it into motion or
mechanical work
; for example, many
protein
-based molecular motors harness the chemical
free energy
released by the
hydrolysis
of ATP in order to perform mechanical work.
[10]
In terms of energetic efficiency, this type of motor can be superior to currently available man-made motors.
Richard Feynman
theorized about the future of
nanomedicine
. He wrote about the idea of a
medical
use for
biological machines
. Feynman and
Albert Hibbs
suggested that certain repair machines might one day be reduced in size to the point that it would be possible to (as Feynman put it) "
swallow the doctor
". The idea was discussed in Feynman's 1959 essay "
There's Plenty of Room at the Bottom
".
[11]
These biological machines might have applications in
nanomedicine
. For example,
[12]
they could be used to identify and destroy cancer cells.
[13]
[14]
Molecular nanotechnology
is a
speculative
subfield of nanotechnology regarding the possibility of engineering
molecular assemblers
, biological machines which could re-order matter at a molecular or atomic scale. Nanomedicine would make use of these
nanorobots
, introduced into the body, to repair or detect damages and infections. Molecular nanotechnology is highly theoretical, seeking to anticipate what inventions nanotechnology might yield and to propose an agenda for future inquiry. The proposed elements of molecular nanotechnology, such as molecular assemblers and nanorobots are far beyond current capabilities.
[15]
[16]
Protein folding
[
edit
]
 Constituent amino-acids can be analyzed to predict secondary, tertiary and quaternary protein structure.
Constituent amino-acids can be analyzed to predict secondary, tertiary and quaternary protein structure.
Protein folding is the
physical process
by which a
protein
chain acquires its
native
3-dimensional
structure, a
conformation
that is usually biologically functional, in an expeditious and reproducible manner. It is the physical process by which a
polypeptide
folds into its characteristic and functional
three-dimensional structure
from a
random coil
.
[17]
Each protein exists as an unfolded polypeptide or random coil when
translated
from a sequence of mRNA to a linear chain of
amino acids
. This polypeptide lacks any stable (long-lasting) three-dimensional structure (the left hand side of the first figure). As the polypeptide chain is being synthesized by a
ribosome
, the linear chain begins to fold into its three-dimensional structure. Folding begins to occur even during the translation of the polypeptide chain. Amino acids interact with each other to produce a well-defined three-dimensional structure, the folded protein (the right-hand side of the figure), known as the
native state
. The resulting three-dimensional structure is determined by the amino acid sequence or primary structure (
Anfinsen's dogma
).
[18]
Protein structure determination
[
edit
]
As the three-dimensional structure of proteins brings with it an understanding of its function and biological context, there is great effort placed in observing the structures of proteins. X-ray crystallography was the primary method used in the 20th century to solve the structures of proteins in their crystalline form. Ever since the early 2000s,
cryogenic electron microscopy
has been used to solve the structures of proteins closer to their native state, as well as observing cellular structures.
[19]
Protein structure prediction
[
edit
]
Protein structure prediction is the inference of the three-dimensional structure of a protein from its
amino acid
sequence?that is, the prediction of its
folding
and its
secondary
and
tertiary structure
from its
primary structure
. Structure prediction is fundamentally different from the inverse problem of
protein design
. Protein structure prediction is one of the most important goals pursued by
bioinformatics
and
theoretical chemistry
; it is highly important in
medicine
, in
drug design
,
biotechnology
and in the design of novel
enzymes
). Every two years, the performance of current methods is assessed in the
CASP
experiment (Critical Assessment of Techniques for Protein Structure Prediction). A continuous evaluation of protein structure prediction web servers is performed by the community project
CAMEO3D
.
The challenge in predicting protein structures is that there lacks a physical model that can fully predict protein tertiary structures from their amino acid sequence. This problem is known as the
de novo
protein structure prediction
problem and is one of the great problems of modern science.
[20]
AlphaFold
, an
artificial intelligence
program, is able to accurately predict the structures of proteins with genetic
homology
to other proteins that have been previously solved.
[21]
Though, this is not a solution to the
de novo
problem, as it relies on a database of prior data which results in it always being biased.
[22]
The solution to the
de novo
protein structure prediction problem must be a purely physical model that will simulate the protein folding in its native environment, resulting in the
in silico
observation of protein structures and dynamics that were never previously observed.
[23]
Spectroscopy
[
edit
]
Spectroscopic techniques like NMR,
spin label
electron spin resonance
,
Raman spectroscopy
,
infrared spectroscopy
,
circular dichroism
, and so on have been widely used to understand structural dynamics of important
biomolecules
and
intermolecular interactions
.
See also
[
edit
]
References
[
edit
]
- ^
"The goal of Molecular Biophysics"
.
Program in Molecular Biophysics
. Johns Hopkins University. Archived from
the original
on 15 March 2012.
- ^
Jackson MB (2006).
Molecular and Cellular Biophysics
. New York: Cambridge University Press.
ISBN
978-1-139-44724-9
.
- ^
Ouzounis CA (2012).
"Rise and demise of bioinformatics? Promise and progress"
.
PLOS Computational Biology
.
8
(4): e1002487.
Bibcode
:
2012PLSCB...8E2487O
.
doi
:
10.1371/journal.pcbi.1002487
.
PMC
3343106
.
PMID
22570600
.
- ^
"NIH working definition of bioinformatics and computational biology"
(PDF)
. Biomedical Information Science and Technology Initiative. 17 July 2000. Archived from
the original
(PDF)
on 5 September 2012
. Retrieved
18 August
2012
.
- ^
"About the CCMB"
.
Center for Computational Molecular Biology
. Providence RI: Brown University
. Retrieved
18 August
2012
.
- ^
Zimmerberg J (April 2006).
"Membrane biophysics"
.
Current Biology
.
16
(8): R272?R276.
Bibcode
:
2006CBio...16.R272Z
.
doi
:
10.1016/j.cub.2006.03.050
.
PMID
16631568
.
- ^
Satir P, Christensen ST (June 2008).
"Structure and function of mammalian cilia"
.
Histochemistry and Cell Biology
.
129
(6): 687?693.
doi
:
10.1007/s00418-008-0416-9
.
PMC
2386530
.
PMID
18365235
. 1432-119X.
- ^
Kinbara K, Aida T (April 2005). "Toward intelligent molecular machines: directed motions of biological and artificial molecules and assemblies".
Chemical Reviews
.
105
(4): 1377?1400.
doi
:
10.1021/cr030071r
.
PMID
15826015
.
- ^
Bu Z, Callaway DJ (2011). "Proteins MOVE! Protein dynamics and long-range allostery in cell signaling".
Protein Structure and Diseases
. Advances in Protein Chemistry and Structural Biology. Vol. 83. pp. 163?221.
doi
:
10.1016/B978-0-12-381262-9.00005-7
.
ISBN
9780123812629
.
PMID
21570668
.
- ^
Bustamante C, Chemla YR, Forde NR, Izhaky D (2004). "Mechanical processes in biochemistry".
Annual Review of Biochemistry
.
73
: 705?748.
doi
:
10.1146/annurev.biochem.72.121801.161542
.
PMID
15189157
.
- ^
Feynman RP (December 1959).
"There's Plenty of Room at the Bottom"
. Archived from
the original
on 2010-02-11
. Retrieved
2017-01-01
.
- ^
Amrute-Nayak M, Diensthuber RP, Steffen W, Kathmann D, Hartmann FK, Fedorov R, et al. (2010). "Targeted optimization of a protein nanomachine for operation in biohybrid devices".
Angewandte Chemie
.
49
(2): 312?316.
Bibcode
:
2010AngCh.122..322A
.
doi
:
10.1002/ange.200905200
.
PMID
19921669
.
- ^
Patel GM, Patel GC, Patel RB, Patel JK, Patel M (February 2006). "Nanorobot: a versatile tool in nanomedicine".
Journal of Drug Targeting
.
14
(2): 63?67.
doi
:
10.1080/10611860600612862
.
PMID
16608733
.
S2CID
25551052
.
- ^
Balasubramanian S, Kagan D, Hu CM, Campuzano S, Lobo-Castanon MJ, Lim N, et al. (April 2011).
"Micromachine-enabled capture and isolation of cancer cells in complex media"
.
Angewandte Chemie
.
50
(18): 4161?4164.
doi
:
10.1002/anie.201100115
.
PMC
3119711
.
PMID
21472835
.
- ^
Freitas Jr RA (December 2005). Kasabov N, Havukkala I (eds.).
"Current Status of Nanomedicine and Medical Nanorobotics"
(PDF)
.
Journal of Computational and Theoretical Nanoscience
.
2
(4): 471.
Bibcode
:
2005JCTN....2..471K
.
doi
:
10.1166/jctn.2005.001
.
- ^
Freitas Jr RA, Merkle RC (2006).
"Nanofactory Collaboration"
.
Molecular Assembler
.
- ^
Alberts B
, Johnson A, Lewis J, Raff M, Roberts K, Walters P (2002).
"The Shape and Structure of Proteins"
.
Molecular Biology of the Cell; Fourth Edition
. New York and London: Garland Science.
ISBN
978-0-8153-3218-3
.
- ^
Anfinsen CB (July 1972).
"The formation and stabilization of protein structure"
.
The Biochemical Journal
.
128
(4): 737?749.
doi
:
10.1042/bj1280737
.
PMC
1173893
.
PMID
4565129
.
- ^
Curry S (July 2015).
"Structural Biology: A Century-long Journey into an Unseen World"
.
Interdisciplinary Science Reviews
.
40
(3): 308?328.
Bibcode
:
2015ISRv...40..308C
.
doi
:
10.1179/0308018815Z.000000000120
.
PMC
4697198
.
PMID
26740732
.
- ^
"So Much More to Know"
.
Science
.
309
(5731): 78?102. 2005-07-01.
doi
:
10.1126/science.309.5731.78b
.
ISSN
0036-8075
.
- ^
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. (2021-08-26).
"Highly accurate protein structure prediction with AlphaFold"
.
Nature
.
596
(7873): 583?589.
Bibcode
:
2021Natur.596..583J
.
doi
:
10.1038/s41586-021-03819-2
.
ISSN
0028-0836
.
PMC
8371605
.
PMID
34265844
.
- ^
Terwilliger TC, Liebschner D, Croll TI, Williams CJ, McCoy AJ, Poon BK, et al. (2023-11-30).
"AlphaFold predictions are valuable hypotheses and accelerate but do not replace experimental structure determination"
.
Nature Methods
.
21
(1): 110?116.
doi
:
10.1038/s41592-023-02087-4
.
ISSN
1548-7091
.
PMC
10776388
.
PMID
38036854
.
- ^
Dill KA, Ozkan SB, Weikl TR, Chodera JD, Voelz VA (2007-06-01).
"The protein folding problem: when will it be solved?"
.
Current Opinion in Structural Biology
. Nucleic acids / Sequences and topology.
17
(3): 342?346.
doi
:
10.1016/j.sbi.2007.06.001
.
ISSN
0959-440X
.
PMID
17572080
.